Selection and characterization of pre-mRNA splicing enhancers: identification of novel SR protein-specific enhancer sequences
- PMID: 10022858
- PMCID: PMC83964
- DOI: 10.1128/MCB.19.3.1705
Selection and characterization of pre-mRNA splicing enhancers: identification of novel SR protein-specific enhancer sequences
Abstract
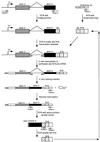
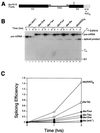
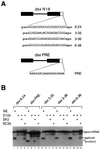
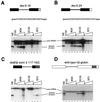
Splicing enhancers are RNA sequences required for accurate splice site recognition and the control of alternative splicing. In this study, we used an in vitro selection procedure to identify and characterize novel RNA sequences capable of functioning as pre-mRNA splicing enhancers. Randomized 18-nucleotide RNA sequences were inserted downstream from a Drosophila doublesex pre-mRNA enhancer-dependent splicing substrate. Functional splicing enhancers were then selected by multiple rounds of in vitro splicing in nuclear extracts, reverse transcription, and selective PCR amplification of the spliced products. Characterization of the selected splicing enhancers revealed a highly heterogeneous population of sequences, but we identified six classes of recurring degenerate sequence motifs five to seven nucleotides in length including novel splicing enhancer sequence motifs. Analysis of selected splicing enhancer elements and other enhancers in S100 complementation assays led to the identification of individual enhancers capable of being activated by specific serine/arginine (SR)-rich splicing factors (SC35, 9G8, and SF2/ASF). In addition, a potent splicing enhancer sequence isolated in the selection specifically binds a 20-kDa SR protein. This enhancer sequence has a high level of sequence homology with a recently identified RNA-protein adduct that can be immunoprecipitated with an SRp20-specific antibody. We conclude that distinct classes of selected enhancers are activated by specific SR proteins, but there is considerable sequence degeneracy within each class. The results presented here, in conjunction with previous studies, reveal a remarkably broad spectrum of RNA sequences capable of binding specific SR proteins and/or functioning as SR-specific splicing enhancers.
Figures

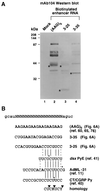

References
-
- Abmayr S M, Workman J L. Preparation of nuclear and cytoplasmic extracts from mammalian cells. In: Ausubel F, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. Vol. 2. New York, N.Y: Greene Publishing Associates and Wiley-Interscience; 1987. pp. 12.1.1–12.1.9.
-
- Adams M D, Rudner D Z, Rio D C. Biochemistry and regulation of pre-mRNA splicing. Curr Opin Cell Biol. 1996;8:331–339. - PubMed
-
- Amrein H, Gorman M, Nöthiger R. The sex-determining gene tra-2 of Drosophila encodes a putative RNA binding protein. Cell. 1988;55:1025–1035. - PubMed
-
- Amrein H, Hedley M L, Maniatis T. The role of specific protein-RNA and protein-protein interactions in positive and negative control of pre-mRNA splicing by Transformer-2. Cell. 1994;76:735–746. - PubMed
-
- Baker B S. Sex in flies: the splice of life. Nature. 1989;340:521–524. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
