Processing of endogenous pre-mRNAs in association with SC-35 domains is gene specific
- PMID: 10037785
- PMCID: PMC2132926
- DOI: 10.1083/jcb.144.4.617
Processing of endogenous pre-mRNAs in association with SC-35 domains is gene specific
Abstract
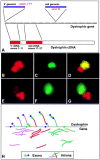
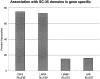
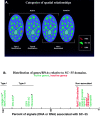
Analysis of six endogenous pre-mRNAs demonstrates that localization at the periphery or within splicing factor-rich (SC-35) domains is not restricted to a few unusually abundant pre-mRNAs, but is apparently a more common paradigm of many protein-coding genes. Different genes are preferentially transcribed and their RNAs processed in different compartments relative to SC-35 domains. These differences do not simply correlate with the complexity, nuclear abundance, or position within overall nuclear space. The distribution of spliceosome assembly factor SC-35 did not simply mirror the distribution of individual pre-mRNAs, but rather suggested that individual domains contain both specific pre-mRNA(s) as well as excess splicing factors. This is consistent with a multifunctional compartment, to which some gene loci and their RNAs have access and others do not. Despite similar molar abundance in muscle fiber nuclei, nascent transcript "trees" of highly complex dystrophin RNA are cotranscriptionally spliced outside of SC-35 domains, whereas posttranscriptional "tracks" of more mature myosin heavy chain transcripts overlap domains. Further analyses supported that endogenous pre-mRNAs exhibit distinct structural organization that may reflect not only the expression and complexity of the gene, but also constraints of its chromosomal context and kinetics of its RNA metabolism.
Figures






Similar articles
-
Nonrandom gene organization: structural arrangements of specific pre-mRNA transcription and splicing with SC-35 domains.J Cell Biol. 1995 Dec;131(6 Pt 2):1635-47. doi: 10.1083/jcb.131.6.1635. J Cell Biol. 1995. PMID: 8557734 Free PMC article.
-
Nuclear pre-mRNA compartmentalization: trafficking of released transcripts to splicing factor reservoirs.Mol Biol Cell. 2000 Feb;11(2):497-510. doi: 10.1091/mbc.11.2.497. Mol Biol Cell. 2000. PMID: 10679009 Free PMC article.
-
Tracking COL1A1 RNA in osteogenesis imperfecta. splice-defective transcripts initiate transport from the gene but are retained within the SC35 domain.J Cell Biol. 2000 Aug 7;150(3):417-32. doi: 10.1083/jcb.150.3.417. J Cell Biol. 2000. PMID: 10931857 Free PMC article.
-
Cross talk between spliceosome and microprocessor defines the fate of pre-mRNA.Wiley Interdiscip Rev RNA. 2014 Sep-Oct;5(5):647-58. doi: 10.1002/wrna.1236. Epub 2014 Apr 30. Wiley Interdiscip Rev RNA. 2014. PMID: 24788135 Review.
-
Plant serine/arginine-rich proteins: roles in precursor messenger RNA splicing, plant development, and stress responses.Wiley Interdiscip Rev RNA. 2011 Nov-Dec;2(6):875-89. doi: 10.1002/wrna.98. Epub 2011 Jul 15. Wiley Interdiscip Rev RNA. 2011. PMID: 21766458 Review.
Cited by
-
FBI-1 can stimulate HIV-1 Tat activity and is targeted to a novel subnuclear domain that includes the Tat-P-TEFb-containing nuclear speckles.Mol Biol Cell. 2002 Mar;13(3):915-29. doi: 10.1091/mbc.01-08-0383. Mol Biol Cell. 2002. PMID: 11907272 Free PMC article.
-
Functional dissection of NEAT1 using genome editing reveals substantial localization of the NEAT1_1 isoform outside paraspeckles.RNA. 2017 Jun;23(6):872-881. doi: 10.1261/rna.059477.116. Epub 2017 Mar 21. RNA. 2017. PMID: 28325845 Free PMC article.
-
Hematopoietic cell transplantation provides an immune-tolerant platform for myoblast transplantation in dystrophic dogs.Mol Ther. 2008 Jul;16(7):1340-6. doi: 10.1038/mt.2008.102. Epub 2008 May 27. Mol Ther. 2008. PMID: 18500253 Free PMC article.
-
Analysis of allele-specific RNA transcription in FSHD by RNA-DNA FISH in single myonuclei.Eur J Hum Genet. 2010 Apr;18(4):448-56. doi: 10.1038/ejhg.2009.183. Epub 2009 Nov 4. Eur J Hum Genet. 2010. PMID: 19888305 Free PMC article.
-
Gene positioning.Cold Spring Harb Perspect Biol. 2010 Jun;2(6):a000588. doi: 10.1101/cshperspect.a000588. Epub 2010 May 19. Cold Spring Harb Perspect Biol. 2010. PMID: 20484389 Free PMC article. Review.
References
-
- Carter KC, Bowman D, Carrington W, Fogarty K, McNeil JA, Fay FS, Lawrence JB. A three-dimensional view of precursor messenger RNA metabolism within the mammalian nucleus. Science. 1993;259:1330–1335. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

