Characterization of the exopolygalacturonate lyase PelX of Erwinia chrysanthemi 3937
- PMID: 10049400
- PMCID: PMC93558
- DOI: 10.1128/JB.181.5.1652-1663.1999
Characterization of the exopolygalacturonate lyase PelX of Erwinia chrysanthemi 3937
Abstract
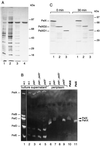
Erwinia chrysanthemi 3937 secretes several pectinolytic enzymes, among which eight isoenzymes of pectate lyases with an endo-cleaving mode (PelA, PelB, PelC, PelD, PelE, PelI, PelL, and PelZ) have been identified. Two exo-cleaving enzymes, the exopolygalacturonate lyase, PelX, and an exo-poly-alpha-D-galacturonosidase, PehX, have been previously identified in other E. chrysanthemi strains. Using a genomic bank of a 3937 mutant with the major pel genes deleted, we cloned a pectinase gene identified as pelX, encoding the exopolygalacturonate lyase. The deduced amino acid sequence of the 3937 PelX is very similar to the PelX of another E. chrysanthemi strain, EC16, except in the 43 C-terminal amino acids. PelX also has homology to the endo-pectate lyase PelL of E. chrysanthemi but has a N-terminal extension of 324 residues. The transcription of pelX, analyzed by gene fusions, is dependent on several environmental conditions. It is induced by pectic catabolic products and affected by growth phase, oxygen limitation, nitrogen starvation, and catabolite repression. Regulation of pelX expression is dependent on the KdgR repressor, which controls almost all the steps of pectin catabolism, and on the global activator of sugar catabolism, cyclic AMP receptor protein. In contrast, PecS and PecT, two repressors of the transcription of most pectate lyase genes, are not involved in pelX expression. The pelX mutant displayed reduced pathogenicity on chicory leaves, but its virulence on potato tubers or Saintpaulia ionantha plants did not appear to be affected. The purified PelX protein has no maceration activity on plant tissues. Tetragalacturonate is the best substrate of PelX, but PelX also has good activity on longer oligomers. Therefore, the estimated number of binding subsites for PelX is 4, extending from subsites -2 to +2. PelX and PehX were shown to be localized in the periplasm of E. chrysanthemi 3937. PelX catalyzed the formation of unsaturated digalacturonates by attack from the reducing end of the substrate, while PehX released digalacturonates by attack from the nonreducing end of the substrate. Thus, the two types of exo-degrading enzymes appeared complementary in the degradation of pectic polymers, since they act on both extremities of the polymeric chain.
Figures


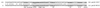





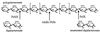
References
-
- Bardonnet N, Blanco C. uidA antibiotic resistance cassettes for insertion mutagenesis, gene fusion and genetic constructions. FEMS Microbiol Lett. 1992;93:243–248. - PubMed
-
- Barras F, Van Gijsegem F, Chatterjee A K. Extracellular enzymes and pathogenesis of soft-rot Erwinia. Annu Rev Phytopathol. 1994;32:201–234.
-
- Beaulieu C, Boccara M, Van Gijsegem F. Pathogenic behaviour of pectinase-defective Erwinia chrysanthemi mutants on different plants. Mol Plant-Microbe Interact. 1993;6:197–202.
-
- Beer, S. Personal communication.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

