Human PEX19: cDNA cloning by functional complementation, mutation analysis in a patient with Zellweger syndrome, and potential role in peroxisomal membrane assembly
- PMID: 10051604
- PMCID: PMC26746
- DOI: 10.1073/pnas.96.5.2116
Human PEX19: cDNA cloning by functional complementation, mutation analysis in a patient with Zellweger syndrome, and potential role in peroxisomal membrane assembly
Abstract
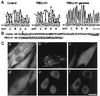
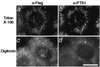
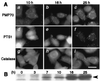
At least 11 complementation groups (CGs) have been identified for the peroxisome biogenesis disorders (PBDs) such as Zellweger syndrome, for which seven pathogenic genes have been elucidated. We have isolated a human PEX19 cDNA (HsPEX19) by functional complementation of peroxisome deficiency of a mutant Chinese hamster ovary cell line, ZP119, defective in import of both matrix and membrane proteins. This cDNA encodes a hydrophilic protein (Pex19p) comprising 299 amino acids, with a prenylation motif, CAAX box, at the C terminus. Farnesylated Pex19p is partly, if not all, anchored in the peroxisomal membrane, exposing its N-terminal part to the cytosol. A stable transformant of ZP119 with HsPEX19 was morphologically and biochemically restored for peroxisome biogenesis. HsPEX19 expression also restored peroxisomal protein import in fibroblasts from a patient (PBDJ-01) with Zellweger syndrome of CG-J. This patient (PBDJ-01) possessed a homozygous, inactivating mutation: a 1-base insertion, A764, in a codon for Met255, resulted in a frameshift, inducing a 24-aa sequence entirely distinct from normal Pex19p. These results demonstrate that PEX19 is the causative gene for CG-J PBD and suggest that the C-terminal part, including the CAAX homology box, is required for the biological function of Pex19p. Moreover, Pex19p is apparently involved at the initial stage in peroxisome membrane assembly, before the import of matrix protein.
Figures




Similar articles
-
Human PEX1 cloned by functional complementation on a CHO cell mutant is responsible for peroxisome-deficient Zellweger syndrome of complementation group I.Proc Natl Acad Sci U S A. 1998 Apr 14;95(8):4350-5. doi: 10.1073/pnas.95.8.4350. Proc Natl Acad Sci U S A. 1998. PMID: 9539740 Free PMC article.
-
Mutation in PEX16 is causal in the peroxisome-deficient Zellweger syndrome of complementation group D.Am J Hum Genet. 1998 Dec;63(6):1622-30. doi: 10.1086/302161. Am J Hum Genet. 1998. PMID: 9837814 Free PMC article.
-
PEX12, the pathogenic gene of group III Zellweger syndrome: cDNA cloning by functional complementation on a CHO cell mutant, patient analysis, and characterization of PEX12p.Mol Cell Biol. 1998 Jul;18(7):4324-36. doi: 10.1128/MCB.18.7.4324. Mol Cell Biol. 1998. PMID: 9632816 Free PMC article.
-
Peroxisome biogenesis disorders: molecular basis for impaired peroxisomal membrane assembly: in metabolic functions and biogenesis of peroxisomes in health and disease.Biochim Biophys Acta. 2012 Sep;1822(9):1337-42. doi: 10.1016/j.bbadis.2012.06.004. Epub 2012 Jun 13. Biochim Biophys Acta. 2012. PMID: 22705440 Review.
-
[Molecular biology of peroxisome biogenesis].Nihon Rinsho. 1993 Sep;51(9):2336-42. Nihon Rinsho. 1993. PMID: 8411711 Review. Japanese.
Cited by
-
PEX13 is mutated in complementation group 13 of the peroxisome-biogenesis disorders.Am J Hum Genet. 1999 Sep;65(3):621-34. doi: 10.1086/302534. Am J Hum Genet. 1999. PMID: 10441568 Free PMC article.
-
Pex17p is required for import of both peroxisome membrane and lumenal proteins and interacts with Pex19p and the peroxisome targeting signal-receptor docking complex in Pichia pastoris.Mol Biol Cell. 1999 Dec;10(12):4005-19. doi: 10.1091/mbc.10.12.4005. Mol Biol Cell. 1999. PMID: 10588639 Free PMC article.
-
Interaction of a farnesylated protein with renal type IIa Na/Pi co-transporter in response to parathyroid hormone and dietary phosphate.Biochem J. 2004 Feb 1;377(Pt 3):607-16. doi: 10.1042/BJ20031223. Biochem J. 2004. PMID: 14558883 Free PMC article.
-
A Humanized Yeast System for Evaluating the Protein Prenylation of a Wide Range of Human and Viral CaaX Sequences.bioRxiv [Preprint]. 2023 Sep 20:2023.09.19.558494. doi: 10.1101/2023.09.19.558494. bioRxiv. 2023. Update in: Dis Model Mech. 2024 May 1;17(5):dmm050516. doi: 10.1242/dmm.050516. PMID: 37786692 Free PMC article. Updated. Preprint.
-
The peroxisomal membrane protein import receptor Pex3p is directly transported to peroxisomes by a novel Pex19p- and Pex16p-dependent pathway.J Cell Biol. 2008 Dec 29;183(7):1275-86. doi: 10.1083/jcb.200806062. J Cell Biol. 2008. PMID: 19114594 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

