Genetic interactions between KAR7/SEC71, KAR8/JEM1, KAR5, and KAR2 during nuclear fusion in Saccharomyces cerevisiae
- PMID: 10069807
- PMCID: PMC25191
- DOI: 10.1091/mbc.10.3.609
Genetic interactions between KAR7/SEC71, KAR8/JEM1, KAR5, and KAR2 during nuclear fusion in Saccharomyces cerevisiae
Abstract
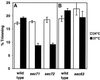
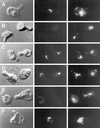
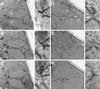
During mating of Saccharomyces cerevisiae, two nuclei fuse to produce a single diploid nucleus. Two genes, KAR7 and KAR8, were previously identified by mutations that cause defects in nuclear membrane fusion. KAR7 is allelic to SEC71, a gene involved in protein translocation into the endoplasmic reticulum. Two other translocation mutants, sec63-1 and sec72Delta, also exhibited moderate karyogamy defects. Membranes from kar7/sec71Delta and sec72Delta, but not sec63-1, exhibited reduced membrane fusion in vitro, but only at elevated temperatures. Genetic interactions between kar7 and kar5 mutations were suggestive of protein-protein interactions. Moreover, in sec71 mutants, Kar5p was absent from the SPB and was not detected by Western blot or immunoprecipitation of pulse-labeled protein. KAR8 is allelic to JEMI, encoding an endoplasmic reticulum resident DnaJ protein required for nuclear fusion. Overexpression of KAR8/JEM1 (but not SEC63) strongly suppressed the mating defect of kar2-1, suggesting that Kar2p interacts with Kar8/Jem1p for nuclear fusion. Electron microscopy analysis of kar8 mutant zygotes revealed a nuclear fusion defect different from kar2, kar5, and kar7/sec71 mutants. Analysis of double mutants suggested that Kar5p acts before Kar8/Jem1p. We propose the existence of a nuclear envelope fusion chaperone complex in which Kar2p, Kar5p, and Kar8/Jem1p are key components and Sec71p and Sec72p play auxiliary roles.
Figures




References
-
- Beh CT. Functions of the Yeast Endoplasmic Reticulum. Ph.D. Thesis. Princeton, NJ: Princeton University; 1996.
-
- Byers B. Multiple roles of the spindle pole bodies in the life cycle of Saccharomyces cerevisiae. In: von Wittstein D, Friis J, Kielland-Brandt M, Stenderup A, editors. Molecular Genetics of Yeast. Copenhagen: Munksgaard; 1981. pp. 119–131.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

