Overexpression of a novel Arabidopsis gene related to putative zinc-transporter genes from animals can lead to enhanced zinc resistance and accumulation
- PMID: 10069843
- PMCID: PMC32086
- DOI: 10.1104/pp.119.3.1047
Overexpression of a novel Arabidopsis gene related to putative zinc-transporter genes from animals can lead to enhanced zinc resistance and accumulation
Abstract
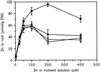
We describe the isolation of an Arabidopsis gene that is closely related to the animal ZnT genes (Zn transporter). The protein encoded by the ZAT (Zn transporter of Arabidopsis thaliana) gene has 398 amino acid residues and is predicted to have six membrane-spanning domains. To obtain evidence for the postulated function of the Arabidopsis gene, transgenic plants with the ZAT coding sequence under control of the cauliflower mosaic virus 35S promoter were analyzed. Plants obtained with ZAT in the sense orientation exhibited enhanced Zn resistance and strongly increased Zn content in the roots under high Zn exposure. Antisense mRNA-producing plants were viable, with a wild-type level of Zn resistance and content, like plants expressing a truncated coding sequence lacking the C-terminal cytoplasmic domain of the protein. The availability of ZAT can lead to a better understanding of the mechanism of Zn homeostasis and resistance in plants.
Figures






References
-
- Conklin D, Culbertson MR, Kung C. Interactions between gene products involved in divalent cation transport in Saccharomyces cerevisiae. Mol Gen Genet. 1994;244:303–311. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

