Analysis of elements involved in pseudoknot-dependent expression and regulation of the repA gene of an IncL/M plasmid
- PMID: 10074073
- PMCID: PMC93579
- DOI: 10.1128/JB.181.6.1811-1819.1999
Analysis of elements involved in pseudoknot-dependent expression and regulation of the repA gene of an IncL/M plasmid
Abstract
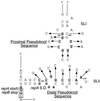
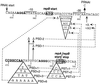
Replication of the IncL/M plasmid pMU604 is controlled by a small antisense RNA molecule (RNAI), which, by inhibiting the formation of an RNA pseudoknot, regulates translation of the replication initiator protein, RepA. Efficient translation of the repA mRNA was shown to require the translation and correct termination of the leader peptide, RepB, and the formation of the pseudoknot. Although the pseudoknot was essential for the expression of repA, its presence was shown to interfere with the translation of repB. The requirement for pseudoknot formation could in large part be obviated by improving the ribosome binding region of repA, either by replacing the GUG start codon by AUG or by increasing the spacing between the start codon and the Shine-Dalgarno sequence (SD). The spacing between the distal pseudoknot sequence and the repA SD was shown to be suboptimal for maximal expression of repA.
Figures

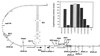
References
-
- Asano K, Kato A, Moriwaki H, Hama C, Shiba K, Mizobuchi K. Positive and negative regulations of plasmid ColIb-P9 repZ gene expression at the translational level. J Biol Chem. 1991;266:3774–3781. - PubMed
-
- Asano K, Mizobuchi K. An RNA pseudoknot as the molecular switch for translation of the repZ gene encoding the replication initiator of IncIα plasmid ColIb-P9. J Biol Chem. 1998;273:11815–11825. - PubMed
-
- Asano K, Moriwaki H, Mizobuchi K. An induced mRNA secondary structure enhances repZ translation in plasmid ColIb-P9. J Biol Chem. 1991;266:24549–24556. - PubMed
-
- Asano K, Niimi T, Yokoyama S, Mizobuchi K. Structural basis for binding of the plasmid ColIb-P9 antisense Inc RNA to its target RNA with the 5′-rUUGGCG-3′ motif in the loop sequence. J Biol Chem. 1998;273:11826–11838. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

