Novel protein kinases Ark1p and Prk1p associate with and regulate the cortical actin cytoskeleton in budding yeast
- PMID: 10087264
- PMCID: PMC2150571
- DOI: 10.1083/jcb.144.6.1203
Novel protein kinases Ark1p and Prk1p associate with and regulate the cortical actin cytoskeleton in budding yeast
Abstract
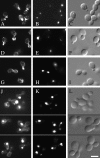
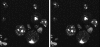
Ark1p (actin regulating kinase 1) was identified as a yeast protein that binds to Sla2p, an evolutionarily conserved cortical actin cytoskeleton protein. Ark1p and a second yeast protein, Prk1p, contain NH2-terminal kinase domains that are 70% identical. Together with six other putative kinases from a number of organisms, these proteins define a new protein kinase family that we have named the Ark family. Lack of both Ark1p and Prk1p resulted in the formation of large cytoplasmic actin clumps and severe defects in cell growth. These defects were rescued by wild-type, but not by kinase-dead versions of the proteins. Elevated levels of either Ark1p or Prk1p caused a number of actin and cell morphological defects that were not observed when the kinase-dead versions were overexpressed instead. Ark1p and Prk1p were shown to localize to actin cortical patches, making these two kinases the first signaling proteins demonstrated to be patch components. These results suggest that Ark1p and Prk1p may be downstream effectors of signaling pathways that control actin patch organization and function. Furthermore, results of double-mutant analyses suggest that Ark1p and Prk1p function in overlapping but distinct pathways that regulate the cortical actin cytoskeleton.
Figures








References
-
- Adams AE, Botstein D, Drubin DG. A yeast actin-binding protein is encoded by SAC6, a gene found by suppression of an actin mutation. Science. 1989;243:231–233. - PubMed
-
- Adams AE, Botstein D, Drubin DG. Requirement of yeast fimbrin for actin organization and morphogenesis in vivo. Nature. 1991;354:404–408. - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Arber S, Barbayannis FA, Hanser H, Schneider C, Stanyon CA, Bernard O, Caroni P. Regulation of actin dynamics through phosphorylation of cofilin by LIM-kinase. Nature. 1998;393:805–809. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

