SMRTe, a silencing mediator for retinoid and thyroid hormone receptors-extended isoform that is more related to the nuclear receptor corepressor
- PMID: 10097068
- PMCID: PMC22325
- DOI: 10.1073/pnas.96.7.3519
SMRTe, a silencing mediator for retinoid and thyroid hormone receptors-extended isoform that is more related to the nuclear receptor corepressor
Abstract
SMRT (silencing mediator for retinoid and thyroid hormone receptors) and N-CoR (nuclear receptor copressor) mediate transcriptional repression of important regulators that are involved in many signaling pathways. SMRT and N-CoR are related proteins that form complexes with mSin3A/B and histone deacetylases to induce local chromatin condensation and transcriptional repression. However, SMRT is substantially smaller than N-CoR, lacking an N-terminal domain of approximately 1,000 aa that are present in N-CoR. Here, we report the identification of SMRT-extended (SMRTe), which contains an N-terminal sequence that shows striking similarity with N-CoR. As in N-CoR, this SMRTe-N-terminal domain also represses basal transcription. We find that SMRTe expression is regulated during cell cycle progression and SMRTe transcripts are present in many embryonic tissues. These data redefine a structurally and functionally more related nuclear receptor corepressor family and suggest an additional role for SMRTe in the regulation of cycle-specific gene expression in diverse signaling pathways.
Figures


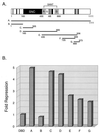


References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

