Single-pair fluorescence resonance energy transfer on freely diffusing molecules: observation of Förster distance dependence and subpopulations
- PMID: 10097095
- PMCID: PMC22352
- DOI: 10.1073/pnas.96.7.3670
Single-pair fluorescence resonance energy transfer on freely diffusing molecules: observation of Förster distance dependence and subpopulations
Abstract
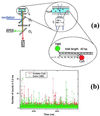
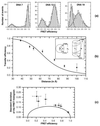
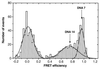
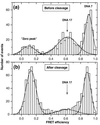
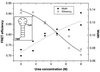
Photon bursts from single diffusing donor-acceptor labeled macromolecules were used to measure intramolecular distances and identify subpopulations of freely diffusing macromolecules in a heterogeneous ensemble. By using DNA as a rigid spacer, a series of constructs with varying intramolecular donor-acceptor spacings were used to measure the mean and distribution width of fluorescence resonance energy transfer (FRET) efficiencies as a function of distance. The mean single-pair FRET efficiencies qualitatively follow the distance dependence predicted by Förster theory. Possible contributions to the widths of the FRET efficiency distributions are discussed, and potential applications in the study of biopolymer conformational dynamics are suggested. The ability to measure intramolecular (and intermolecular) distances for single molecules implies the ability to distinguish and monitor subpopulations of molecules in a mixture with different distances or conformational states. This is demonstrated by monitoring substrate and product subpopulations before and after a restriction endonuclease cleavage reaction. Distance measurements at single-molecule resolution also should facilitate the study of complex reactions such as biopolymer folding. To this end, the denaturation of a DNA hairpin was examined by using single-pair FRET.
Figures
References
-
- Basché T, Moerner W E, Orrit M, Wild U P, editors. Single-Molecule Optical Detection, Imaging, and Spectroscopy. Cambridge: Wiley; 1997.
-
- Nie S M, Zare R N. Annu Rev Biophys Biomol Struct. 1997;26:567–596. - PubMed
-
- Xie X S, Trautman J K. Annu Rev Phys Chem. 1998;49:441–480. - PubMed
-
- Lu H P, Xun L, Xie X S. Science. 1998;282:1877–1882. - PubMed
-
- Shera E B, Seitzinger N K, Davis L M, Keller R A, Soper S A. Chem Phys Lett. 1990;174:553–557.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

