The FEZ1 gene at chromosome 8p22 encodes a leucine-zipper protein, and its expression is altered in multiple human tumors
- PMID: 10097140
- PMCID: PMC22397
- DOI: 10.1073/pnas.96.7.3928
The FEZ1 gene at chromosome 8p22 encodes a leucine-zipper protein, and its expression is altered in multiple human tumors
Abstract
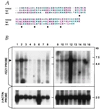
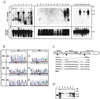
Alterations of human chromosome 8p occur frequently in many tumors. We identified a 1.5-Mb common region of allelic loss on 8p22 by allelotype analysis. cDNA selection allowed isolation of several genes, including FEZ1. The predicted Fez1 protein contained a leucine-zipper region with similarity to the DNA-binding domain of the cAMP-responsive activating-transcription factor 5. RNA blot analysis revealed that FEZ1 gene expression was undetectable in more than 60% of epithelial tumors. Mutations were found in primary esophageal cancers and in a prostate cancer cell line. Transcript analysis from several FEZ1-expressing tumors revealed truncated mRNAs, including a frameshift. Alteration and inactivation of the FEZ1 gene may play a role in various human tumors.
Figures

References
-
- Cavenee W, Dryja T P, Phillips R A, Benedict W F, Godbout R, Gallie B L, Murphree A L, Strong L C, White R L. Nature (London) 1983;305:779–784. - PubMed
-
- Lasko D, Cavenee W. Annu Rev Genet. 1991;25:281–314. - PubMed
-
- Weinberg R. Science. 1991;254:1138–1146. - PubMed
-
- Nowell P. Adv Cancer Res. 1993;62:1–17. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

