Apg7p/Cvt2p is required for the cytoplasm-to-vacuole targeting, macroautophagy, and peroxisome degradation pathways
- PMID: 10233148
- PMCID: PMC25275
- DOI: 10.1091/mbc.10.5.1337
Apg7p/Cvt2p is required for the cytoplasm-to-vacuole targeting, macroautophagy, and peroxisome degradation pathways
Abstract
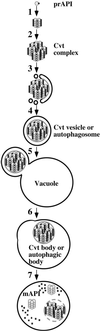
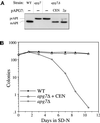
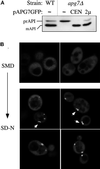
Proper functioning of organelles necessitates efficient protein targeting to the appropriate subcellular locations. For example, degradation in the fungal vacuole relies on an array of targeting mechanisms for both resident hydrolases and their substrates. The particular processes that are used vary depending on the available nutrients. Under starvation conditions, macroautophagy is the primary method by which bulk cytosol is sequestered into autophagic vesicles (autophagosomes) destined for this organelle. Molecular genetic, morphological, and biochemical evidence indicates that macroautophagy shares much of the same cellular machinery as a biosynthetic pathway for the delivery of the vacuolar hydrolase, aminopeptidase I, via the cytoplasm-to-vacuole targeting (Cvt) pathway. The machinery required in both pathways includes a novel protein modification system involving the conjugation of two autophagy proteins, Apg12p and Apg5p. The conjugation reaction was demonstrated to be dependent on Apg7p, which shares homology with the E1 family of ubiquitin-activating enzymes. In this study, we demonstrate that Apg7p functions at the sequestration step in the formation of Cvt vesicles and autophagosomes. The subcellular localization of Apg7p fused to green fluorescent protein (GFP) indicates that a subpopulation of Apg7pGFP becomes membrane associated in an Apg12p-dependent manner. Subcellular fractionation experiments also indicate that a portion of the Apg7p pool is pelletable under starvation conditions. Finally, we demonstrate that the Pichia pastoris homologue Gsa7p that is required for peroxisome degradation is functionally similar to Apg7p, indicating that this novel conjugation system may represent a general nonclassical targeting mechanism that is conserved across species.
Figures





Similar articles
-
Membrane recruitment of Aut7p in the autophagy and cytoplasm to vacuole targeting pathways requires Aut1p, Aut2p, and the autophagy conjugation complex.J Cell Biol. 2001 Jan 8;152(1):51-64. doi: 10.1083/jcb.152.1.51. J Cell Biol. 2001. PMID: 11149920 Free PMC article.
-
Apg7p/Cvt2p: A novel protein-activating enzyme essential for autophagy.Mol Biol Cell. 1999 May;10(5):1367-79. doi: 10.1091/mbc.10.5.1367. Mol Biol Cell. 1999. PMID: 10233150 Free PMC article.
-
Apg5p functions in the sequestration step in the cytoplasm-to-vacuole targeting and macroautophagy pathways.Mol Biol Cell. 2000 Mar;11(3):969-82. doi: 10.1091/mbc.11.3.969. Mol Biol Cell. 2000. PMID: 10712513 Free PMC article.
-
Transport of proteins to the yeast vacuole: autophagy, cytoplasm-to-vacuole targeting, and role of the vacuole in degradation.Semin Cell Dev Biol. 2000 Jun;11(3):173-9. doi: 10.1006/scdb.2000.0163. Semin Cell Dev Biol. 2000. PMID: 10906274 Review.
-
Autophagy, cytoplasm-to-vacuole targeting pathway, and pexophagy in yeast and mammalian cells.Annu Rev Biochem. 2000;69:303-42. doi: 10.1146/annurev.biochem.69.1.303. Annu Rev Biochem. 2000. PMID: 10966461 Review.
Cited by
-
Membrane recruitment of Aut7p in the autophagy and cytoplasm to vacuole targeting pathways requires Aut1p, Aut2p, and the autophagy conjugation complex.J Cell Biol. 2001 Jan 8;152(1):51-64. doi: 10.1083/jcb.152.1.51. J Cell Biol. 2001. PMID: 11149920 Free PMC article.
-
Vacuolar localization of oligomeric alpha-mannosidase requires the cytoplasm to vacuole targeting and autophagy pathway components in Saccharomyces cerevisiae.J Biol Chem. 2001 Jun 8;276(23):20491-8. doi: 10.1074/jbc.M101150200. Epub 2001 Mar 22. J Biol Chem. 2001. PMID: 11264288 Free PMC article.
-
Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells.J Cell Biol. 2001 Feb 19;152(4):657-68. doi: 10.1083/jcb.152.4.657. J Cell Biol. 2001. PMID: 11266458 Free PMC article.
-
The molecular machinery of autophagy: unanswered questions.J Cell Sci. 2005 Jan 1;118(Pt 1):7-18. doi: 10.1242/jcs.01620. J Cell Sci. 2005. PMID: 15615779 Free PMC article.
-
An overview of autophagy: morphology, mechanism, and regulation.Antioxid Redox Signal. 2014 Jan 20;20(3):460-73. doi: 10.1089/ars.2013.5371. Epub 2013 Aug 2. Antioxid Redox Signal. 2014. PMID: 23725295 Free PMC article. Review.
References
-
- Baba M, Osumi M, Ohsumi Y. Analysis of the membrane structures involved in autophagy in yeast by freeze-replica method. Cell Struct Funct. 1995;20:465–471. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

