Differential expression of two novel members of the tomato ethylene-receptor family
- PMID: 10318694
- PMCID: PMC59248
- DOI: 10.1104/pp.120.1.165
Differential expression of two novel members of the tomato ethylene-receptor family
Abstract
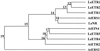
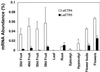
The phytohormone ethylene regulates many aspects of plant growth, development, and environmental responses. Much of the developmental regulation of ethylene responses in tomato (Lycopersicon esculentum) occurs at the level of hormone sensitivity. In an effort to understand the regulation of ethylene responses, we isolated and characterized tomato genes with sequence similarity to the Arabidopsis ETR1 (ethylene response 1) ethylene receptor. Previously, we isolated three genes that exhibit high similarity to ETR1 and to each other. Here we report the isolation of two additional genes, LeETR4 and LeETR5, that are only 42% and 40% identical to ETR1, respectively. Although the amino acids known to be involved in ethylene binding are conserved, LeETR5 lacks the histidine within the kinase domain that is predicted to be phosphorylated. This suggests that histidine kinase activity is not necessary for an ethylene response, because mutated forms of both LeETR4 and LeETR5 confer dominant ethylene insensitivity in transgenic Arabidopsis plants. Expression analysis indicates that LeETR4 accounts for most of the putative ethylene-receptor mRNA present in reproductive tissues, but, like LeETR5, it is less abundant in vegetative tissues. Taken together, ethylene perception in tomato is potentially quite complex, with at least five structurally divergent, putative receptor family members exhibiting significant variation in expression levels throughout development.
Figures


Similar articles
-
The tomato ethylene receptors NR and LeETR4 are negative regulators of ethylene response and exhibit functional compensation within a multigene family.Proc Natl Acad Sci U S A. 2000 May 9;97(10):5663-8. doi: 10.1073/pnas.090550597. Proc Natl Acad Sci U S A. 2000. PMID: 10792050 Free PMC article.
-
Differential regulation of the tomato ETR gene family throughout plant development.Plant J. 1998 Jul;15(2):243-52. doi: 10.1046/j.1365-313x.1998.00202.x. Plant J. 1998. PMID: 9721682
-
Evidence that CTR1-mediated ethylene signal transduction in tomato is encoded by a multigene family whose members display distinct regulatory features.Plant Mol Biol. 2004 Feb;54(3):387-404. doi: 10.1023/B:PLAN.0000036371.30528.26. Plant Mol Biol. 2004. PMID: 15284494
-
The ethylene-receptor family from Arabidopsis: structure and function.Philos Trans R Soc Lond B Biol Sci. 1998 Sep 29;353(1374):1405-12. doi: 10.1098/rstb.1998.0295. Philos Trans R Soc Lond B Biol Sci. 1998. PMID: 9800203 Free PMC article. Review.
-
Control of ethylene-mediated processes in tomato at the level of receptors.J Exp Bot. 2002 Oct;53(377):2057-63. doi: 10.1093/jxb/erf062. J Exp Bot. 2002. PMID: 12324529 Review.
Cited by
-
Cloning and characterisation of two CTR1-like genes in Cucurbita pepo: regulation of their expression during male and female flower development.Sex Plant Reprod. 2010 Dec;23(4):301-13. doi: 10.1007/s00497-010-0140-1. Sex Plant Reprod. 2010. PMID: 20390430
-
Energy conservation and dissipation in mitochondria isolated from developing tomato fruit of ethylene-defective mutants failing normal ripening: the effect of ethephon, a chemical precursor of ethylene.J Bioenerg Biomembr. 2003 Apr;35(2):157-68. doi: 10.1023/a:1023750204310. J Bioenerg Biomembr. 2003. PMID: 12887014
-
Functional analysis of the Arlequin mutant corroborates the essential role of the Arlequin/TAGL1 gene during reproductive development of tomato.PLoS One. 2010 Dec 23;5(12):e14427. doi: 10.1371/journal.pone.0014427. PLoS One. 2010. PMID: 21203447 Free PMC article.
-
Role of ethylene receptors during senescence and ripening in horticultural crops.Plant Signal Behav. 2012 Jul;7(7):827-46. doi: 10.4161/psb.20321. Epub 2012 Jul 1. Plant Signal Behav. 2012. PMID: 22751331 Free PMC article. Review.
-
Evidence of the functional role of the ethylene receptor genes SlETR4 and SlETR5 in ethylene signal transduction in tomato.Mol Genet Genomics. 2019 Apr;294(2):301-313. doi: 10.1007/s00438-018-1505-7. Epub 2018 Oct 31. Mol Genet Genomics. 2019. PMID: 30382349
References
-
- Abeles FB, Morgan PW, Saltveit ME (1992). Ethylene in Plant Biology, Ed 2. Academic Press, San Diego, CA, pp 264–296
-
- Bechtold N, Ellis J, Pelletier G. In planta Agrobacterium-mediated gene transfer by infiltration of adult Arabidopsis plants. CR Acad Sci Paris. 1993;316:1194–1199.
-
- Bleecker AB, Estelle MA, Somerville C, Kende H. Insensitivity to ethylene conferred by a dominant mutation in Arabidopsis. Science. 1988;241:1086–1089. - PubMed
-
- Chang C, Kwok SF, Bleecker AB, Meyerowitz EM. Arabidopsis ethylene-response gene ETR1: similarity of products to two-component regulators. Science. 1993;262:539–544. - PubMed
-
- Chao Q, Rothenberg M, Solano R, Roman G, Terzaghi W, Ecker J. Activation of the ethylene gas response pathway in Arabidopsis by the nuclear protein ETHYLENE-INSENSITIVE3 and related proteins. Cell. 1997;89:1133–1144. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

