Replication mechanism and sequence analysis of the replicon of pAW63, a conjugative plasmid from Bacillus thuringiensis
- PMID: 10322022
- PMCID: PMC93776
- DOI: 10.1128/JB.181.10.3193-3200.1999
Replication mechanism and sequence analysis of the replicon of pAW63, a conjugative plasmid from Bacillus thuringiensis
Abstract
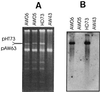
A 5.8-kb fragment of the large conjugative plasmid pAW63 from Bacillus thuringiensis subsp. kurstaki HD73 containing all the information for autonomous replication was cloned and sequenced. By deletion analysis, the pAW63 replicon was reduced to a 4.1-kb fragment harboring four open reading frames (ORFs). Rep63A (513 amino acids [aa]), encoded by the largest ORF, displayed strong similarity (40% identity) to the replication proteins from plasmids pAMbeta1, pIP501, and pSM19035, indicating that the pAW63 replicon belongs to the pAMbeta1 family of gram-positive theta-replicating plasmids. This was confirmed by the facts that no single-stranded DNA replication intermediates could be detected and that replication was found to be dependent on host-gene-encoded DNA polymerase I. An 85-bp region downstream of Rep63A was also shown to have strong similarity to the origins of replication of pAMbeta1 and pIP501, and it is suggested that this region contains the bona fide pAW63 ori. The protein encoded by the second large ORF, Rep63B (308 aa), was shown to display similarity to RepB (34% identity over 281 aa) and PrgP (32% identity over 310 aa), involved in copy control of the Enterococcus faecalis plasmids pAD1 and pCF10, respectively. No significant similarity to known proteins or DNA sequences could be detected for the two smallest ORFs. However, the location, size, hydrophilicity, and orientation of ORF6 (107 codons) were analogous to those features of the putative genes repC and prgO, which encode stability functions on plasmids pAD1 and pCF10, respectively. The cloned replicon of plasmid pAW63 was stably maintained in Bacillus subtilis and B. thuringiensis and displayed incompatibility with the native pAW63. Hybridization experiments using the cloned replicon as a probe showed that pAW63 has similarity to large plasmids from other B. thuringiensis subsp. kurstaki strains and to a strain of B. thuringiensis subsp. alesti.
Figures

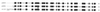


References
-
- Andrup L, Damgaard J, Wassermann K, Boe L, Madsen S M, Hansen F G. Complete nucleotide sequence of the Bacillus thuringiensis subsp. israelensis plasmid pTX14-3 and its correlation with biological properties. Plasmid. 1994;31:72–88. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. Brooklyn, N.Y: John Wiley & Sons, Inc.; 1995.
-
- Baum J A, Gonzalez J M. Mode of replication, size and distribution of naturally occurring plasmids in Bacillus thuringiensis. FEMS Microbiol Lett. 1992;96:143–148. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

