A TATA-binding protein mutant defective for TFIID complex formation in vivo
- PMID: 10330135
- PMCID: PMC104354
- DOI: 10.1128/MCB.19.6.3951
A TATA-binding protein mutant defective for TFIID complex formation in vivo
Abstract
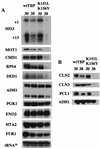
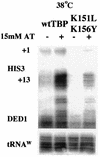
Using an intragenic complementation screen, we have identified a temperature-sensitive TATA-binding protein (TBP) mutant (K151L, K156Y) that is defective for interaction with certain yeast TBP-associated factors (TAFs) at the restrictive temperature. The K151L,K156Y mutant appears to be functional for RNA polymerase I (Pol I) and Pol III transcription, and it is capable of supporting Gal4-activated and Gcn4-activated transcription by Pol II. However, transcription from certain TATA-containing and TATA-less Pol II promoters is reduced at the restrictive temperature. Immunoprecipitation analysis of extracts prepared after culturing cells at the restrictive temperature for 1 h indicates that the K151L,K156Y derivative is severely compromised in its ability to interact with TAF130, TAF90, TAF68/61, and TAF25 while remaining functional for interaction with TAF60 and TAF30. Thus, a TBP mutant that is compromised in its ability to form TFIID can support the response to Gcn4 but is defective for transcription from specific promoters in vivo.
Figures




References
-
- Apone L M, Virbasius C-A, Holstege F C, Wang J, Young R A, Green M R. Broad, but not universal, transcriptional requirement for yTAFII17, a histone H3-like TAFII present in TFIID and SAGA. Mol Cell. 1998;2:653–661. - PubMed
-
- Apone L M, Virbasius C A, Reese J C, Green M R. Yeast TAFII90 is required for cell-cycle progression through G2/M but not for general transcription activation. Genes Dev. 1996;10:2368–2380. - PubMed
-
- Burley S K, Roeder R G. Biochemistry and structural biology of transcription factor IID (TFIID) Annu Rev Biochem. 1996;65:769–799. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
