Ku antigen-DNA conformation determines the activation of DNA-dependent protein kinase and DNA sequence-directed repression of mouse mammary tumor virus transcription
- PMID: 10330147
- PMCID: PMC104366
- DOI: 10.1128/MCB.19.6.4065
Ku antigen-DNA conformation determines the activation of DNA-dependent protein kinase and DNA sequence-directed repression of mouse mammary tumor virus transcription
Abstract
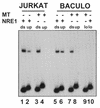
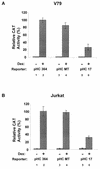
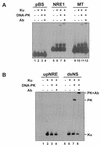
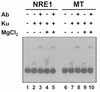
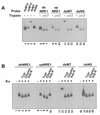
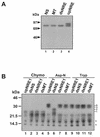
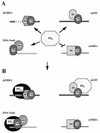
Mouse mammary tumor virus (MMTV) transcription is repressed by DNA-dependent protein kinase (DNA-PK) through a DNA sequence element, NRE1, in the viral long terminal repeat that is a sequence-specific DNA binding site for the Ku antigen subunit of the kinase. While Ku is an essential component of the active kinase, how the catalytic subunit of DNA-PK (DNA-PKcs) is regulated through its association with Ku is only beginning to be understood. We report that activation of DNA-PKcs and the repression of MMTV transcription from NRE1 are dependent upon Ku conformation, the manipulation of DNA structure by Ku, and the contact of Ku80 with DNA. Truncation of one copy of the overlapping direct repeat that comprises NRE1 abrogated the repression of MMTV transcription by Ku-DNA-PKcs. Remarkably, the truncated element was recognized by Ku-DNA-PKcs with affinity similar to that of the full-length element but was unable to promote the activation of DNA-PKcs. Analysis of Ku-DNA-PKcs interactions with DNA ends, double- and single-stranded forms of NRE1, and the truncated NRE1 element revealed striking differences in Ku conformation that differentially affected the recruitment of DNA-PKcs and the activation of kinase activity.
Figures





Similar articles
-
Sequence-specific binding of Ku autoantigen to single-stranded DNA.J Biol Chem. 1998 Aug 14;273(33):20810-9. doi: 10.1074/jbc.273.33.20810. J Biol Chem. 1998. PMID: 9694826
-
Sequence-specific DNA binding by Ku autoantigen and its effects on transcription.Nature. 1996 Mar 21;380(6571):265-8. doi: 10.1038/380265a0. Nature. 1996. PMID: 8637578
-
Ku represses the HIV-1 transcription: identification of a putative Ku binding site homologous to the mouse mammary tumor virus NRE1 sequence in the HIV-1 long terminal repeat.J Biol Chem. 2002 Feb 15;277(7):4918-24. doi: 10.1074/jbc.M110830200. Epub 2001 Nov 30. J Biol Chem. 2002. PMID: 11733502
-
Interaction of Ku protein and DNA-dependent protein kinase catalytic subunit with nucleic acids.Nucleic Acids Res. 1998 Apr 1;26(7):1551-9. doi: 10.1093/nar/26.7.1551. Nucleic Acids Res. 1998. PMID: 9512523 Free PMC article. Review.
-
Ku, a DNA repair protein with multiple cellular functions?Mutat Res. 1999 May 14;434(1):3-15. doi: 10.1016/s0921-8777(99)00006-3. Mutat Res. 1999. PMID: 10377944 Review.
Cited by
-
Host Proteins Ku and HMGA1 As Participants of HIV-1 Transcription.Acta Naturae. 2016 Jan-Mar;8(1):34-47. Acta Naturae. 2016. PMID: 27099783 Free PMC article.
-
Impact of the Ku complex on HIV-1 expression and latency.PLoS One. 2013 Jul 29;8(7):e69691. doi: 10.1371/journal.pone.0069691. Print 2013. PLoS One. 2013. PMID: 23922776 Free PMC article.
-
DNA-PK contributes to the phosphorylation of AIRE: importance in transcriptional activity.Biochim Biophys Acta. 2008 Jan;1783(1):74-83. doi: 10.1016/j.bbamcr.2007.09.003. Epub 2007 Oct 4. Biochim Biophys Acta. 2008. PMID: 17997173 Free PMC article.
-
Computational studies on full-length Ku70 with DNA duplexes: base interactions and a helical path.J Mol Model. 2012 May;18(5):1935-49. doi: 10.1007/s00894-011-1220-3. Epub 2011 Aug 26. J Mol Model. 2012. PMID: 21870196
-
Subnuclear localization of Ku protein: functional association with RNA polymerase II elongation sites.Mol Cell Biol. 2002 Nov;22(22):8088-99. doi: 10.1128/MCB.22.22.8088-8099.2002. Mol Cell Biol. 2002. PMID: 12391174 Free PMC article.
References
-
- Anderson C W, Lees-Miller S P. The nuclear serine/threonine protein kinase DNA-PK. Crit Rev Eukaryot Gene Expr. 1992;2:283–314. - PubMed
-
- Archer T K, Lefebvre P, Wolford R G, Hager G L. Transcription factor loading on the MMTV promoter: a bimodal mechanism for promoter activation. Science. 1992;255:1573–1576. - PubMed
-
- Beato M. Chromatin structure and the regulation of gene expression: remodeling at the MMTV promoter. J Mol Med. 1996;74:711–724. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
