KAP-1 corepressor protein interacts and colocalizes with heterochromatic and euchromatic HP1 proteins: a potential role for Krüppel-associated box-zinc finger proteins in heterochromatin-mediated gene silencing
- PMID: 10330177
- PMCID: PMC104396
- DOI: 10.1128/MCB.19.6.4366
KAP-1 corepressor protein interacts and colocalizes with heterochromatic and euchromatic HP1 proteins: a potential role for Krüppel-associated box-zinc finger proteins in heterochromatin-mediated gene silencing
Abstract
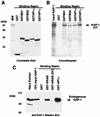
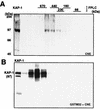
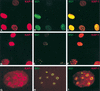
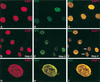
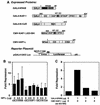
Krüppel-associated box (KRAB) domains are present in approximately one-third of all human zinc finger proteins (ZFPs) and are potent transcriptional repression modules. We have previously cloned a corepressor for the KRAB domain, KAP-1, which is required for KRAB-mediated repression in vivo. To characterize the repression mechanism utilized by KAP-1, we have analyzed the ability of KAP-1 to interact with murine (M31 and M32) and human (HP1alpha and HP1gamma) homologues of the HP1 protein family, a class of nonhistone heterochromatin-associated proteins with a well-established epigenetic gene silencing function in Drosophila. In vitro studies confirmed that KAP-1 is capable of directly interacting with M31 and hHP1alpha, which are normally found in centromeric heterochromatin, as well as M32 and hHP1gamma, both of which are found in euchromatin. Mapping of the region in KAP-1 required for HP1 interaction showed that amino acid substitutions which abolish HP1 binding in vitro reduce KAP-1 mediated repression in vivo. We observed colocalization of KAP-1 with M31 and M32 in interphase nuclei, lending support to the biochemical evidence that M31 and M32 directly interact with KAP-1. The colocalization of KAP-1 with M31 is sometimes found in subnuclear territories of potential pericentromeric heterochromatin, whereas colocalization of KAP-1 and M32 occurs in punctate euchromatic domains throughout the nucleus. This work suggests a mechanism for the recruitment of HP1-like gene products by the KRAB-ZFP-KAP-1 complex to specific loci within the genome through formation of heterochromatin-like complexes that silence gene activity. We speculate that gene-specific repression may be a consequence of the formation of such complexes, ultimately leading to silenced genes in newly formed heterochromatic chromosomal environments.
Figures


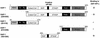



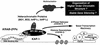
References
-
- Albagli O, Dhordain P, Deweindt C, Lecocq G, Leprince D. The BTB/POZ domain: a new protein-protein interaction motif common to DNA- and actin-binding proteins. Cell Growth Differ. 1995;6:1193–1198. - PubMed
-
- Auble D T, Hansen K E, Mueller C G, Lane W S, Thorner J, Hahn S. Mot1, a global repressor of RNA polymerase II transcription, inhibits TBP binding to DNA by an ATP-dependent mechanism. Genes Dev. 1994;8:1920–1934. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
