Vertical distribution of methanogens in the anoxic sediment of Rotsee (Switzerland)
- PMID: 10347020
- PMCID: PMC91355
- DOI: 10.1128/AEM.65.6.2402-2408.1999
Vertical distribution of methanogens in the anoxic sediment of Rotsee (Switzerland)
Abstract
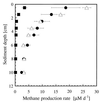
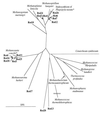
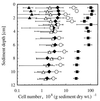
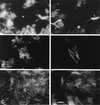
Anoxic sediments from Rotsee (Switzerland) were analyzed for the presence and diversity of methanogens by using molecular tools and for methanogenic activity by using radiotracer techniques, in addition to the measurement of chemical profiles. After PCR-assisted sequence retrieval of the 16S rRNA genes (16S rDNA) from the anoxic sediment of Rotsee, cloning, and sequencing, a phylogenetic analysis identified two clusters of sequences and four separated clones. The sequences in cluster 1 grouped with those of Methanosaeta spp., whereas the sequences in cluster 2 comprised the methanogenic endosymbiont of Plagiopyla nasuta. Discriminative oligonucleotide probes were constructed against both clusters and two of the separated clones. These probes were used subsequently for the analysis of indigenous methanogens in a core of the sediment, in addition to domain-specific probes against members of the domains Bacteria and Archaea and the fluorescent stain 4', 6-diamidino-2-phenylindole (DAPI), by fluorescent in situ hybridization. After DAPI staining, the highest microbial density was obtained in the upper sediment layer; this density decreased with depth from (1.01 +/- 0.25) x 10(10) to (2.62 +/- 0.58) x 10(10) cells per g of sediment (dry weight). This zone corresponded to that of highest metabolic activity, as indicated by the ammonia, alkalinity, and pH profiles, whereas the methane profile was constant. Probes Eub338 and Arch915 detected on average 16 and 6% of the DAPI-stained cells as members of the domains Bacteria and Archaea, respectively. Probe Rotcl1 identified on average 4% of the DAPI-stained cells as Methanosaeta spp., which were present throughout the whole core. In contrast, probe Rotcl2 identified only 0.7% of the DAPI-stained cells as relatives of the methanogenic endosymbiont of P. nasuta, which was present exclusively in the upper 2 cm of the sediment. Probes Rotp13 and Rotp17 did not detect any cells. The spatial distribution of the two methanogenic populations corresponded well to the methane production rates determined by incubation with either [14C]acetate or [14C]bicarbonate. Methanogenesis from acetate accounted for almost all of the total methane production, which concurs with the predominance of acetoclastic Methanosaeta spp. that represented on average 91% of the archaeal population. Significant hydrogenotrophic methanogenesis was found only in the organically enriched upper 2 cm of the sediment, where the probably hydrogenotrophic relatives of the methanogenic endosymbiont of P. nasuta, accounting on average for 7% of the archaeal population, were also detected.
Figures

References
-
- Ammann A A, Rüttimann T B. Simultaneous determination of small organic and inorganic anions in environmental water samples by ion-exchange chromatography. J Chromatogr. 1995;706:259–269.
-
- Beveridge T J, Harris B J, Patel G B, Sprott G D. Cell division and filament splitting in Methanothrix concilii. Can J Microbiol. 1986;33:725–732.
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

