Importance of a 5' stem-loop for longevity of papA mRNA in Escherichia coli
- PMID: 10348874
- PMCID: PMC93829
- DOI: 10.1128/JB.181.11.3587-3590.1999
Importance of a 5' stem-loop for longevity of papA mRNA in Escherichia coli
Abstract
High-level expression of the major pilus subunit (PapA) of uropathogenic strains of Escherichia coli results in part from the unusually long lifetime of the mRNA that encodes this protein. Here we report that the longevity of papA mRNA derives in large measure from the protection afforded by its 5' untranslated region. This papA RNA segment can prolong the lifetime of an otherwise short-lived mRNA to which it is fused. In vivo alkylation studies indicate that, in its natural milieu, the papA message begins with a stem-loop structure. This stem-loop is important for the stabilizing effect of the papA 5' untranslated region, as evidenced by the significant acceleration in papA mRNA decay that results from its removal.
Figures

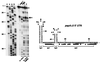

Similar articles
-
An evolutionarily conserved RNA stem-loop functions as a sensor that directs feedback regulation of RNase E gene expression.Genes Dev. 2000 May 15;14(10):1249-60. Genes Dev. 2000. PMID: 10817759 Free PMC article.
-
A conserved RNA structure (thi box) is involved in regulation of thiamin biosynthetic gene expression in bacteria.Proc Natl Acad Sci U S A. 2001 Aug 14;98(17):9736-41. doi: 10.1073/pnas.161168098. Epub 2001 Jul 24. Proc Natl Acad Sci U S A. 2001. PMID: 11470904 Free PMC article.
-
A 5' stem-loop and ribosome binding but not translation are important for the stability of Bacillus subtilis aprE leader mRNA.Microbiology (Reading). 2002 Jun;148(Pt 6):1795-1803. doi: 10.1099/00221287-148-6-1795. Microbiology (Reading). 2002. PMID: 12055299
-
Translation initiation and the fate of bacterial mRNAs.FEMS Microbiol Rev. 2006 Nov;30(6):967-79. doi: 10.1111/j.1574-6976.2006.00043.x. Epub 2006 Sep 21. FEMS Microbiol Rev. 2006. PMID: 16989654 Review.
-
Measuring the immeasurable.Mol Cell. 2002 Sep;10(3):437-9. doi: 10.1016/s1097-2765(02)00661-5. Mol Cell. 2002. PMID: 12408812 Review.
Cited by
-
Adjacent single-stranded regions mediate processing of tRNA precursors by RNase E direct entry.Nucleic Acids Res. 2014 Apr;42(7):4577-89. doi: 10.1093/nar/gkt1403. Epub 2014 Jan 21. Nucleic Acids Res. 2014. PMID: 24452799 Free PMC article.
-
A VapBC toxin-antitoxin module is a posttranscriptional regulator of metabolic flux in mycobacteria.J Bacteriol. 2012 May;194(9):2189-204. doi: 10.1128/JB.06790-11. Epub 2012 Feb 24. J Bacteriol. 2012. PMID: 22366418 Free PMC article.
-
The Bacillus subtilis late competence operon comE is transcriptionally regulated by yutB and under post-transcription initiation control by comN (yrzD).J Bacteriol. 2009 Feb;191(3):949-58. doi: 10.1128/JB.01429-08. Epub 2008 Nov 21. J Bacteriol. 2009. PMID: 19028902 Free PMC article.
-
Initiation of RNA decay in Escherichia coli by 5' pyrophosphate removal.Mol Cell. 2007 Jul 6;27(1):79-90. doi: 10.1016/j.molcel.2007.05.038. Mol Cell. 2007. PMID: 17612492 Free PMC article.
-
Regulation of clpQ⁺Y⁺ (hslV⁺U⁺) gene expression in Escherichia coli.Open Microbiol J. 2009;3:29-39. doi: 10.2174/1874285800903010029. Epub 2009 Mar 17. Open Microbiol J. 2009. PMID: 19440251 Free PMC article.
References
-
- Båga M, Göransson M, Normark S, Uhlin B E. Processed mRNA with differential stability in the regulation of E. coli pilin gene expression. Cell. 1988;52:197–206. - PubMed
-
- Belasco J G, Nilsson G, von Gabain A, Cohen S N. The stability of E. coli gene transcripts is dependent on determinants localized to specific mRNA segments. Cell. 1986;46:245–251. - PubMed
-
- Bouvet P, Belasco J G. Control of RNase E-mediated RNA degradation by 5′-terminal base pairing in E. coli. Nature. 1992;360:488–491. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

