The human G2 checkpoint control protein hRAD9 is a nuclear phosphoprotein that forms complexes with hRAD1 and hHUS1
- PMID: 10359610
- PMCID: PMC25401
- DOI: 10.1091/mbc.10.6.1985
The human G2 checkpoint control protein hRAD9 is a nuclear phosphoprotein that forms complexes with hRAD1 and hHUS1
Abstract
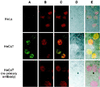
Eukaryotic cells actively block entry into mitosis in the presence of DNA damage or incompletely replicated DNA. This response is mediated by signal transduction cascades called cell cycle checkpoints. We show here that the human checkpoint control protein hRAD9 physically associates with two other checkpoint control proteins, hRAD1 and hHUS1. Furthermore, hRAD1 and hHUS1 themselves interact, analogously to their fission yeast homologues Rad1 and Hus1. We also show that hRAD9 is present in multiple phosphorylation forms in vivo. These phosphorylated forms are present in tissue culture cells that have not been exposed to exogenous sources of DNA damage, but it remains possible that endogenous damage or naturally occurring replication intermediates cause the observed phosphorylation. Finally, we show that hRAD9 is a nuclear protein, indicating that in this signal transduction pathway, hRAD9 is physically proximal to the upstream (DNA damage) signal rather than to the downstream, cytoplasmic, cell cycle machinery.
Figures




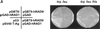
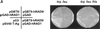

Similar articles
-
Human homologs of Schizosaccharomyces pombe rad1, hus1, and rad9 form a DNA damage-responsive protein complex.J Biol Chem. 1999 Jan 8;274(2):567-70. doi: 10.1074/jbc.274.2.567. J Biol Chem. 1999. PMID: 9872989
-
A role of the C-terminal region of human Rad9 (hRad9) in nuclear transport of the hRad9 checkpoint complex.J Biol Chem. 2002 Jul 12;277(28):25722-7. doi: 10.1074/jbc.M203079200. Epub 2002 May 6. J Biol Chem. 2002. PMID: 11994305
-
Human hRad1 but not hRad9 protects hHus1 from ubiquitin-proteasomal degradation.Oncogene. 2004 Jul 1;23(30):5124-30. doi: 10.1038/sj.onc.1207658. Oncogene. 2004. PMID: 15122316
-
Dial 9-1-1 for DNA damage: the Rad9-Hus1-Rad1 (9-1-1) clamp complex.DNA Repair (Amst). 2004 Aug-Sep;3(8-9):1009-14. doi: 10.1016/j.dnarep.2004.03.032. DNA Repair (Amst). 2004. PMID: 15279787 Review.
-
Cell-cycle responses to DNA damage in G2.Int Rev Cytol. 2003;222:99-140. doi: 10.1016/s0074-7696(02)22013-6. Int Rev Cytol. 2003. PMID: 12503848 Review.
Cited by
-
Characterization of Schizosaccharomyces pombe Hus1: a PCNA-related protein that associates with Rad1 and Rad9.Mol Cell Biol. 2000 Feb;20(4):1254-62. doi: 10.1128/MCB.20.4.1254-1262.2000. Mol Cell Biol. 2000. PMID: 10648611 Free PMC article.
-
Structure-function analysis of fission yeast Hus1-Rad1-Rad9 checkpoint complex.Mol Biol Cell. 2001 Dec;12(12):3744-58. doi: 10.1091/mbc.12.12.3744. Mol Biol Cell. 2001. PMID: 11739777 Free PMC article.
-
Telomeres, histone code, and DNA damage response.Cytogenet Genome Res. 2008;122(3-4):297-307. doi: 10.1159/000167816. Epub 2009 Jan 30. Cytogenet Genome Res. 2008. PMID: 19188699 Free PMC article. Review.
-
Viral transport of DNA damage that mimics a stalled replication fork.J Virol. 2005 Jan;79(1):569-80. doi: 10.1128/JVI.79.1.569-580.2005. J Virol. 2005. PMID: 15596849 Free PMC article.
-
Interaction of apurinic/apyrimidinic endonuclease 2 (Apn2) with Myh1 DNA glycosylase in fission yeast.DNA Repair (Amst). 2014 Mar;15:1-10. doi: 10.1016/j.dnarep.2014.01.001. Epub 2014 Feb 1. DNA Repair (Amst). 2014. PMID: 24559510 Free PMC article.
References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Boddy MN, Furnari B, Mondesert O, Russell P. Replication checkpoint enforced by kinases Cds1 and Chk1. Science. 1998;280:909–912. - PubMed
-
- Cahill DP, Lengauer C, Yu J, Riggins GJ, Willson JK, Markowitz SD, Kinzler KW, Vogelstein B. Mutations of mitotic checkpoint genes in human cancers. Nature. 1998;392:300–303. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

