CPF: an orphan nuclear receptor that regulates liver-specific expression of the human cholesterol 7alpha-hydroxylase gene
- PMID: 10359768
- PMCID: PMC21971
- DOI: 10.1073/pnas.96.12.6660
CPF: an orphan nuclear receptor that regulates liver-specific expression of the human cholesterol 7alpha-hydroxylase gene
Abstract
Cholesterol 7alpha-hydroxylase is the first and rate-limiting enzyme in a pathway through which cholesterol is metabolized to bile acids. The gene encoding cholesterol 7alpha-hydroxylase, CYP7A, is expressed exclusively in the liver. Overexpression of CYP7A in hamsters results in a reduction of serum cholesterol levels, suggesting that the enzyme plays a central role in cholesterol homeostasis. Here, we report the identification of a hepatic-specific transcription factor that binds to the promoter of the human CYP7A gene. We designate this factor CPF, for CYP7A promoter binding factor. Mutation of the CPF binding site within the CYP7A promoter abolished hepatic-specific expression of the gene in transient transfection assays. A cDNA encoding CPF was cloned and identified as a human homolog of the Drosophila orphan nuclear receptor fushi tarazu F1 (Ftz-F1). Cotransfection of a CPF expression plasmid and a CYP7A reporter gene resulted in specific induction of CYP7A-directed transcription. These observations suggest that CPF is a key regulator of human CYP7A gene expression in the liver.
Figures





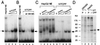


References
-
- Brown M S, Goldstein J L. Science. 1986;232:34–47. - PubMed
-
- Turley S D, Dietschy J M. In: The Liver: Biology and Pathology. Arias I, Popper H, Schachter D, Shafritz D A, editors. New York: Raven; 1982. pp. 467–492.
-
- Goldstein J L, Brown M S. Nature (London) 1990;343:425–430. - PubMed
-
- Brown M S, Goldstein J L. Cell. 1997;89:331–340. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

