Different mutator phenotypes in Mlh1- versus Pms2-deficient mice
- PMID: 10359802
- PMCID: PMC22005
- DOI: 10.1073/pnas.96.12.6850
Different mutator phenotypes in Mlh1- versus Pms2-deficient mice
Abstract
Deficiencies in DNA mismatch repair (MMR) result in increased mutation rates and cancer risk in both humans and mice. Mouse strains homozygous for knockouts of either the Pms2 or Mlh1 MMR gene develop cancer but exhibit very different tumor spectra; only Mlh1(-/-) animals develop intestinal tumors. We carried out a detailed study of the microsatellite mutation spectra in each knockout strain. Five mononucleotide repeat tracts at four different chromosomal locations were studied by using single-molecule PCR or an in vivo forward mutation assay. Three dinucleotide repeat loci also were examined. Surprisingly, the mononucleotide repeat mutation frequency in Mlh1(-/-) mice was 2- to 3-fold higher than in Pms2(-/-) animals. The higher mutation frequency in Mlh1(-/-) mice may be a consequence of some residual DNA repair capacity in the Pms2(-/-) animals. Relevant to this idea, we observed that Pms2(-/-) mice exhibit almost normal levels of Mlh1p, whereas Mlh1(-/-) animals lack both Mlh1p and Pms2p. Comparison between Mlh1(-/-) animals and Mlh1(-/-) and Pms2(-/-) double knockout mice revealed little difference in mutator phenotype, suggesting that Mlh1 nullizygosity is sufficient to inactivate MMR completely. The findings may provide a basis for understanding the greater predisposition to intestinal cancer of Mlh1(-/-) mice. Small differences (2- to 3-fold) in mononucleotide repeat mutation rates may have dramatic effects on tumor development, requiring multiple genetic alterations in coding regions. Alternatively, this strain difference in tumor spectra also may be related to the consequences of the absence of Pms2p compared with the absence of both Pms2p and Mlh1p on as yet little understood cellular processes.
Figures


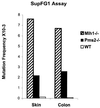

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

