Replication and pathogenicity of primer binding site mutants of SL3-3 murine leukemia viruses
- PMID: 10364369
- PMCID: PMC112678
- DOI: 10.1128/JVI.73.7.6117-6122.1999
Replication and pathogenicity of primer binding site mutants of SL3-3 murine leukemia viruses
Abstract
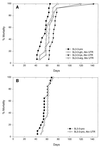
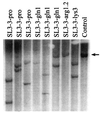
Retroviral reverse transcription is primed by a cellular tRNA molecule annealed to an 18-bp primer binding site sequence. The sequence of the primer binding site coincides with that of a negatively acting cis element that mediates transcriptional silencing of murine leukemia virus (MLV) in undifferentiated embryonic cells. In this study we test whether SL3-3 MLV can replicate stably using tRNA primers other than the cognate tRNAPro and analyze the effect of altering the primer binding site sequence to match the 3' end of tRNA1Gln, tRNA3Lys, or tRNA1,2Arg in a mouse pathogenicity model. Contrary to findings from cell culture studies of primer binding site-modified human immunodeficiency virus type 1 and avian retroviruses, our findings were that SL3-3 MLV may stably and efficiently replicate with tRNA primers other than tRNAPro. Although lymphoma induction of the SL3-3 Lys3 mutant was significantly delayed relative to that of the wild-type virus, molecular tumor analysis indicated that all the primer binding site-modified viruses induce T-cell lymphomas similar to those induced by the wild-type virus in terms of frequencies of genomic rearrangements within the T-cell receptor beta-chain, the immunoglobulin kappa light chain, and the c-myc locus. Whereas none of the mutants were found to revert to tRNAPro primer utilization, in two tumors resulting from the injection of the SL3-3 Lys3 mutant the primer binding site was altered to match that of a new primer species, tRNA1,2Lys. In addition, recombination with endogenous viruses resulting in the generation of recombinant viruses carrying a glutamine primer binding site was detected in the majority of the tumors induced by the SL3-3 Lys3 mutant as well as in two tumors induced by wild-type SL3-3 and the SL3-3 Arg1,2 mutant.
Figures

Similar articles
-
Stability of AML1 (core) site enhancer mutations in T lymphomas induced by attenuated SL3-3 murine leukemia virus mutants.J Virol. 1997 Jul;71(7):5080-7. doi: 10.1128/JVI.71.7.5080-5087.1997. J Virol. 1997. PMID: 9188573 Free PMC article.
-
Increased induction of osteopetrosis, but unaltered lymphomagenicity, by murine leukemia virus SL3-3 after mutation of a nuclear factor 1 site in the enhancer.J Virol. 1999 Dec;73(12):10406-15. doi: 10.1128/JVI.73.12.10406-10415.1999. J Virol. 1999. PMID: 10559359 Free PMC article.
-
Analysis of wild-type and mutant SL3-3 murine leukemia virus insertions in the c-myc promoter during lymphomagenesis reveals target site hot spots, virus-dependent patterns, and frequent error-prone gap repair.J Virol. 2005 Jan;79(1):67-78. doi: 10.1128/JVI.79.1.67-78.2005. J Virol. 2005. PMID: 15596802 Free PMC article.
-
tRNAs as primer of reverse transcriptases.Biochimie. 1995;77(1-2):113-24. doi: 10.1016/0300-9084(96)88114-4. Biochimie. 1995. PMID: 7541250 Review.
-
Viral pathogenesis and immunity within the thymus.Immunol Res. 1998;17(1-2):75-82. doi: 10.1007/BF02786432. Immunol Res. 1998. PMID: 9479569 Review.
Cited by
-
The kissing-loop motif is a preferred site of 5' leader recombination during replication of SL3-3 murine leukemia viruses in mice.J Virol. 1999 Nov;73(11):9614-8. doi: 10.1128/JVI.73.11.9614-9618.1999. J Virol. 1999. PMID: 10516072 Free PMC article.
-
Duplication of the primary encapsidation and dimer linkage region of human immunodeficiency virus type 1 RNA results in the appearance of monomeric RNA in virions.J Virol. 2001 Mar;75(6):2557-65. doi: 10.1128/JVI.75.6.2557-2565.2001. J Virol. 2001. PMID: 11222678 Free PMC article.
-
Selection of functional tRNA primers and primer binding site sequences from a retroviral combinatorial library: identification of new functional tRNA primers in murine leukemia virus replication.Nucleic Acids Res. 2000 Feb 1;28(3):791-9. doi: 10.1093/nar/28.3.791. Nucleic Acids Res. 2000. PMID: 10637332 Free PMC article.
-
Transfer of primer binding site-mutated simian immunodeficiency virus vectors by genetically engineered artificial and hybrid tRNA-like primers.J Virol. 2001 May;75(10):4922-8. doi: 10.1128/JVI.75.10.4922-4928.2001. J Virol. 2001. PMID: 11312366 Free PMC article.
-
Mutation of all Runx (AML1/core) sites in the enhancer of T-lymphomagenic SL3-3 murine leukemia virus unmasks a significant potential for myeloid leukemia induction and favors enhancer evolution toward induction of other disease patterns.J Virol. 2004 Dec;78(23):13216-31. doi: 10.1128/JVI.78.23.13216-13231.2004. J Virol. 2004. PMID: 15542674 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

