Worldwide evaluation of DNA sequencing approaches for identification of drug resistance mutations in the human immunodeficiency virus type 1 reverse transcriptase
- PMID: 10364600
- PMCID: PMC85140
- DOI: 10.1128/JCM.37.7.2291-2296.1999
Worldwide evaluation of DNA sequencing approaches for identification of drug resistance mutations in the human immunodeficiency virus type 1 reverse transcriptase
Abstract
A panel (ENVA-1) of well-defined blinded samples containing wild-type and mutant human immunodeficiency virus type 1 (HIV-1) reverse transcriptase was analyzed by automated DNA sequencing in 23 laboratories worldwide. Drug resistance mutations at codons 41, 215, and 184 were present in the panel samples at different ratios to the wild type. The presence of mutant genotypes was determined qualitatively and quantitatively. All laboratories reported the presence of sequence heterogeneities at codons 41, 215, and 184 in one or more of the panel samples, though not all reported the correct codon genotypes. Two laboratories reported a mutant genotype in samples containing only the wild type, whereas two and three laboratories failed to detect the mutant genotypes at codons 41 and 215, respectively, in a completely mutant DNA population. Mutations present at relative concentrations of 25% of the total DNA population were successfully identified by 13 of 23, 10 of 23, and 16 of 23 labs for codons 41, 215, and 184Val, respectively. For more than 80% of those laboratories that qualitatively detected the presence of a mutation correctly, the estimated wild type/mutant ratio was less than 25% different from the input ratio in those samples containing 25 to 50% or 75% mutant input. This first multicenter study on the quality of DNA sequencing approaches for identifying HIV-1 drug resistance mutations revealed large interlaboratory differences in the quality of the results. The application of these procedures in their current state would in several cases lead to inaccurate or even incorrect diagnostic results. Therefore, proper quality control and standardization are urgently needed.
Figures
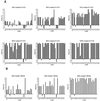
References
-
- Baxter J D, Mayers D L, Wentworth D N, Neaton J D, Merigan T C the CPCRA Study Team for the Terry Beirn Community Programs for Clinical Research on AIDS. Abstracts of the 6th Conference on Retroviruses and Opportunistic Infections. 1999. A pilot study of the short-term effects of antiretroviral management based on plasma genotypic antiretroviral resistance testing (GART) in patients failing antiretorviral therapy, abstr. LB 8.
-
- Boucher C A B, Keulen W, van Bommel T, Nijhuis M, de Jong D, de Jong M D, Schipper P, Back N K T. Human immunodeficiency virus type 1 drug susceptibility determination by using recombinant viruses generated from patient sera tested in a cell-killing assay. Antimicrob Agents Chemother. 1996;40:2404–2409. - PMC - PubMed
-
- Coffin J M. HIV population dynamics in vivo: implications for genetic variation, pathogenesis, and therapy. Science. 1995;267:483–489. - PubMed
-
- Demeter L M, D’Aquila R, Weislow O, Lorenzo E, Erice A, Fitzgibbon J, Shafer R, Richman D, Howard T M, Zhao Y, Fisher E, Huang D, Mayers D, Sylvester S, Arens M, Sannerud K, Rasheed S, Johnson V, Kuritzkes D, Reichelderfer P, Japour A ACTG Sequencing Working Group; AIDS Clinical Trials Group. Interlaboratory concordance of DNA sequence analysis to detect reverse transcriptase mutations in HIV-1 proviral DNA. J Virol Methods. 1998;75:93–104. - PubMed
-
- Durant, J., P. Clevenbergh, P. Halfon, P. Delguidice, S. Porsin, P. Simonet, N. Montagne, E. Dohin, J. M. Schapiro, C. Boucher, and P. Dellamonica. Improving HIV therapy with drug resistance genotyping: The Viradapt Study. Lancet, in press. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

