Mutational analysis of the Streptococcus pneumoniae bimodular class A penicillin-binding proteins
- PMID: 10368166
- PMCID: PMC93869
- DOI: 10.1128/JB.181.12.3852-3856.1999
Mutational analysis of the Streptococcus pneumoniae bimodular class A penicillin-binding proteins
Abstract
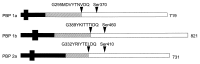
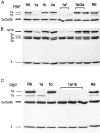
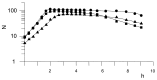
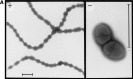
One group of penicillin target enzymes, the class A high-molecular-weight penicillin-binding proteins (PBPs), are bimodular enzymes. In addition to a central penicillin-binding-transpeptidase domain, they contain an N-terminal putative glycosyltransferase domain. Mutations in the genes for each of the three Streptococcus pneumoniae class A PBPs, PBP1a, PBP1b, and PBP2a, were isolated by insertion duplication mutagenesis within the glycosyltransferase domain, documenting that their function is not essential for cellular growth in the laboratory. PBP1b PBP2a and PBP1a PBP1b double mutants could also be isolated, and both showed defects in positioning of the septum. Attempts to obtain a PBP2a PBP1a double mutant failed. All mutants with a disrupted pbp2a gene showed higher sensitivity to moenomycin, an antibiotic known to inhibit PBP-associated glycosyltransferase activity, indicating that PBP2a is the primary target for glycosyltransferase inhibitors in S. pneumoniae.
Figures


References
-
- Adam M, Fraipont C, Rhazi N, Nguyen-Distèche M, Lakaye B, Frère J-M, Devreese B, Van Beeumen J, van Heijenoort Y, Van Heijenoort J, Ghuysen J-M. The bimodular G57-V577 polypeptide chain of the class B penicillin-binding protein 3 of Escherichia colicatalyzes peptide bond formation from thioesters and does not catalyze glycan chain polymerization from lipid II intermediates. J Bacteriol. 1997;179:6005–6009. - PMC - PubMed
-
- Ayala J A, Garrido T, De Pedro M A, Vicente M. Molecular biology of bacterial septation. In: Ghuysen J-M, Hakenbeck R, editors. Bacterial cell wall. Amsterdam, The Netherlands: Elsevier; 1994. pp. 73–101.
-
- Ballester S, Alonso J, Lopez P, Espinoza M. Comparative expression of the pC194 cat gene in Streptococcus pneumoniae, Bacillus subtilis, and Escherichia coli. Gene. 1990;86:71–79. - PubMed
-
- Briese T, Hakenbeck R. Interaction between choline and the N-acetyl-muramyl-L-alanine-amidase of Streptococcus pneumoniae. In: Hakenbeck R, Höltje J-V, Labischinski H, editors. The target of penicillin. Berlin, Germany: Walter de Gruyter & Co.; 1983. pp. 173–178.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

