Characterization and submitochondrial localization of the alpha subunit of the mitochondrial processing peptidase from the aquatic fungus Blastocladiella emersonii
- PMID: 10400583
- PMCID: PMC93927
- DOI: 10.1128/JB.181.14.4257-4265.1999
Characterization and submitochondrial localization of the alpha subunit of the mitochondrial processing peptidase from the aquatic fungus Blastocladiella emersonii
Abstract
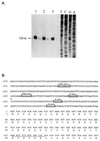
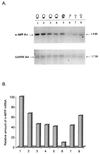
In an effort to investigate the molecular mechanisms responsible for the drastic morphological changes the mitochondria go through during the life cycle of the aquatic fungus Blastocladiella emersonii, the gene encoding the alpha subunit of the mitochondrial processing peptidase (alpha-MPP) was isolated. Nucleotide sequence analysis revealed that the predicted alpha-MPP polypeptide comprises 474 amino acids with a calculated molecular mass of 51,900 Da, presenting a characteristic mitochondrial signal sequence. Northern blot analysis indicated a single 1.4-kb transcript encoding the B. emersonii alpha-MPP, whose levels decrease during sporulation, becoming very low in the zoospore, and increase again during germination. Despite these variations in mRNA concentration, B. emersonii alpha-MPP protein levels do not change significantly during the life cycle of the fungus, as observed in Western blots. Experiments to investigate the submitochondrial localization of B. emersonii alpha-MPP and beta-MPP were also carried out, and the results indicated that both subunits are associated with the mitochondrial inner membrane, possibly as part of the bc1 complex, as described for plants.
Figures
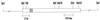



Similar articles
-
Isolation, characterization, and expression of the gene encoding the beta subunit of the mitochondrial processing peptidase from Blastocladiella emersonii.J Bacteriol. 1998 Aug;180(15):3967-72. doi: 10.1128/JB.180.15.3967-3972.1998. J Bacteriol. 1998. PMID: 9683495 Free PMC article.
-
Cloning and structural analysis of the gene for the regulatory subunit of cAMP-dependent protein kinase in Blastocladiella emersonii.J Biol Chem. 1992 Aug 25;267(24):17201-7. J Biol Chem. 1992. PMID: 1512258
-
Cloning and characterization of the gene for the catalytic subunit of cAMP-dependent protein kinase in the aquatic fungus Blastocladiella emersonii.Eur J Biochem. 1994 Jan 15;219(1-2):555-62. doi: 10.1111/j.1432-1033.1994.tb19971.x. Eur J Biochem. 1994. PMID: 8307021
-
Mitochondrial processing peptidases.Biochim Biophys Acta. 2002 Sep 2;1592(1):63-77. doi: 10.1016/s0167-4889(02)00265-3. Biochim Biophys Acta. 2002. PMID: 12191769 Review.
-
Mitochondrial processing peptidase: multiple-site recognition of precursor proteins.Biochem Biophys Res Commun. 1999 Nov 30;265(3):611-6. doi: 10.1006/bbrc.1999.1703. Biochem Biophys Res Commun. 1999. PMID: 10600469 Review.
Cited by
-
Gene discovery and expression profile analysis through sequencing of expressed sequence tags from different developmental stages of the chytridiomycete Blastocladiella emersonii.Eukaryot Cell. 2005 Feb;4(2):455-64. doi: 10.1128/EC.4.2.455-464.2005. Eukaryot Cell. 2005. PMID: 15701807 Free PMC article.
-
Environmental stresses inhibit splicing in the aquatic fungus Blastocladiella emersonii.BMC Microbiol. 2009 Oct 29;9:231. doi: 10.1186/1471-2180-9-231. BMC Microbiol. 2009. PMID: 19874600 Free PMC article.
References
-
- Altschul S F, Gish W, Miller W, Myers E W, Lipman D J. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Avedissian M, Gomes S L. Expression of the groESL operon is cell cycle controlled in Caulobacter crescentus. Mol Microbiol. 1996;19:79–89. - PubMed
-
- Boumans H, Berden J A, Grivell L A. Topological organization of subunits VII and VIII in the ubiquinol-cytochrome c oxidoreductase of Saccharomyces cerevisiae. FEBS Lett. 1996;390:137–141. - PubMed
-
- Bradford M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–251. - PubMed
-
- Braun H P, Schmitz U K. Are the “core” proteins of the mitochondrial bc1 complex evolutionary relics of a processing protease? Trends Biochem Sci. 1995;20:171–175. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

