Characterization of V3 sequence heterogeneity in subtype C human immunodeficiency virus type 1 isolates from Malawi: underrepresentation of X4 variants
- PMID: 10400718
- PMCID: PMC112705
- DOI: 10.1128/JVI.73.8.6271-6281.1999
Characterization of V3 sequence heterogeneity in subtype C human immunodeficiency virus type 1 isolates from Malawi: underrepresentation of X4 variants
Abstract
We have examined the nature of V3 sequence variability among subtype C human immunodeficiency virus type 1 (HIV-1) sequences from plasma-derived viral RNA present in infected men from Malawi. Sequence variability was assessed by direct sequence analysis of the V3 reverse transcription-PCR products, examination of virus populations by a subtype C V3-specific heteroduplex tracking assay (V3-HTA), and selected sequence analysis of molecular clones derived from the PCR products. Sequence variability in V3 among the subtype C viruses was not associated with the presence of basic amino acid substitutions. This observation is in contrast to that for subtype B HIV-1, where sequence variability is associated with such substitutions, and these substitutions are determinants of altered coreceptor usage. Evolutionary variants in subtype C V3 sequences, as defined by the V3-HTA, were not correlated with the CD4 level in the infected person, while such a correlation was found with subtype B V3 sequences. Viruses were isolated from a subset of the subjects; all isolates used CCR5 and not CXCR4 as a coreceptor, and none was able to grow in MT-2 cells, a hallmark of the syncytium-inducing phenotype that is correlated with CXCR4 usage. The overall sequence variability of the subtype C V3 region was no greater than that of the conserved regions of gp120. This limited sequence variability was also a feature of subtype B V3 sequences that do not carry the basic amino acid substitutions associated with altered coreceptor usage. Our results indicate that altered coreceptor usage is rare in subtype C HIV-1 isolates in sub-Saharan Africa and that sequence variability is not a feature of the V3 region of env in the absence of altered coreceptor usage.
Figures

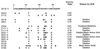


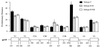
References
-
- Alizon M, Wain-Hobson S, Montagnier L, Sonigo P. Genetic variability of the AIDS virus: nucleotide sequence analysis of two isolates from African patients. Cell. 1986;46:63–74. - PubMed
-
- Alkhatib G, Combadiere C, Broder C C, Feng Y, Kennedy P E, Murphy P M, Berger E A. CC CKR5: a RANTES, MIP-1a, MIP-1b receptor as a fusion cofactor for macrophage-tropic HIV-1. Science. 1996;272:1955–1958. - PubMed
-
- Anzala A O, Ball T B, Rostron T, O’Brien S J, Plummer F A, Rowland-Jones S L University of Nairobi Collaboration for HIV Research. CCR2-64I allele and genotype association with delayed AIDS progression in African women. Lancet. 1998;351:1632–1633. - PubMed
-
- Anzala O A, Nagelkerke N J, Bwayo J J, Holton D, Moses S, Ngugi E N, Ndinya-Achola J O, Plummer F A. Rapid progression to disease in African sex workers with human immunodeficiency virus type 1 infection. J Infect Dis. 1995;171:686–689. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

