Identification of a novel structural protein of arteriviruses
- PMID: 10400725
- PMCID: PMC112712
- DOI: 10.1128/JVI.73.8.6335-6345.1999
Identification of a novel structural protein of arteriviruses
Abstract
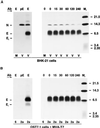
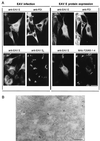
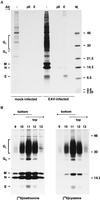
Arteriviruses are positive-stranded RNA viruses with an efficiently organized, polycistronic genome. A short region between the replicase gene and open reading frame (ORF) 2 of the equine arteritis virus (EAV) genome was previously assumed to be untranslated. However, here we report that this segment of the EAV genome contains the 5' part of a novel gene (ORF 2a) which is conserved in all arteriviruses. The 3' part of EAV ORF 2a overlaps with the 5' part of the former ORF 2 (now renamed ORF 2b), which encodes the GS glycoprotein. Both ORF 2a and ORF 2b appear to be expressed from mRNA 2, which thereby constitutes the first proven example of a bicistronic mRNA in arteriviruses. The 67-amino-acid protein encoded by EAV ORF 2a, which we have provisionally named the envelope (E) protein, is very hydrophobic and has a basic C terminus. An E protein-specific antiserum was raised and used to demonstrate the expression of the novel gene in EAV-infected cells. The EAV E protein proved to be very stable, did not form disulfide-linked oligomers, and was not N-glycosylated. Immunofluorescence and immunoelectron microscopy studies showed that the E protein associates with intracellular membranes both in EAV-infected cells and upon independent expression. An analysis of purified EAV particles revealed that the E protein is a structural protein. By using reverse genetics, we demonstrated that both the EAV E and GS proteins are essential for the production of infectious progeny virus.
Figures





Similar articles
-
Characterization of two new structural glycoproteins, GP(3) and GP(4), of equine arteritis virus.J Virol. 2002 Nov;76(21):10829-40. doi: 10.1128/jvi.76.21.10829-10840.2002. J Virol. 2002. PMID: 12368326 Free PMC article.
-
Structural proteins of equine arteritis virus.J Virol. 1992 Nov;66(11):6294-303. doi: 10.1128/JVI.66.11.6294-6303.1992. J Virol. 1992. PMID: 1328669 Free PMC article.
-
Discovery of a small arterivirus gene that overlaps the GP5 coding sequence and is important for virus production.J Gen Virol. 2011 May;92(Pt 5):1097-1106. doi: 10.1099/vir.0.029264-0. Epub 2011 Feb 9. J Gen Virol. 2011. PMID: 21307223 Free PMC article.
-
Experiences with infectious cDNA clones of equine arteritis virus: lessons learned and insights gained.Virology. 2014 Aug;462-463:388-403. doi: 10.1016/j.virol.2014.04.029. Epub 2014 Jun 7. Virology. 2014. PMID: 24913633 Free PMC article. Review.
-
Equine arteritis virus.Vet Microbiol. 2013 Nov 29;167(1-2):93-122. doi: 10.1016/j.vetmic.2013.06.015. Epub 2013 Jul 3. Vet Microbiol. 2013. PMID: 23891306 Free PMC article. Review.
Cited by
-
Research Progress of Porcine Reproductive and Respiratory Syndrome Virus NSP2 Protein.Viruses. 2023 Nov 24;15(12):2310. doi: 10.3390/v15122310. Viruses. 2023. PMID: 38140551 Free PMC article. Review.
-
Expression of the Heterotrimeric GP2/GP3/GP4 Spike of an Arterivirus in Mammalian Cells.Viruses. 2022 Apr 1;14(4):749. doi: 10.3390/v14040749. Viruses. 2022. PMID: 35458479 Free PMC article.
-
Development of a fluorescent-microsphere immunoassay for detection of antibodies specific to equine arteritis virus and comparison with the virus neutralization test.Clin Vaccine Immunol. 2008 Jan;15(1):76-87. doi: 10.1128/CVI.00388-07. Epub 2007 Nov 21. Clin Vaccine Immunol. 2008. PMID: 18032597 Free PMC article.
-
Importance of M-protein C terminus as substrate antigen for serodetection of equine arteritis virus infection.Clin Diagn Lab Immunol. 2002 May;9(3):698-703. doi: 10.1128/cdli.9.3.698-703.2002. Clin Diagn Lab Immunol. 2002. PMID: 11986280 Free PMC article.
-
New insights about the regulation of Nidovirus subgenomic mRNA synthesis.Virology. 2018 Apr;517:38-43. doi: 10.1016/j.virol.2018.01.026. Epub 2018 Feb 21. Virology. 2018. PMID: 29475599 Free PMC article.
References
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons, Inc.; 1987.
-
- Bass B L. RNA editing and hypermutation by adenosine deamination. Trends Biochem Sci. 1997;22:157–162. - PubMed
-
- Bhatnagar R S, Gordon J I. Understanding covalent modifications of proteins by lipids; where cell biology and biophysics mingle. Trends Cell Biol. 1997;7:14–20. - PubMed
-
- Bonifacino J S, Marks M S, Ohno H, Kirchhausen T. Mechanisms of signal-mediated protein sorting in the endocytic and secretory pathways. Proc Assoc Am Phys. 1996;108:285–295. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

