Flanking regulatory sequences of the Tetrahymena R deletion element determine the boundaries of DNA rearrangement
- PMID: 10409752
- PMCID: PMC84415
- DOI: 10.1128/MCB.19.8.5631
Flanking regulatory sequences of the Tetrahymena R deletion element determine the boundaries of DNA rearrangement
Abstract
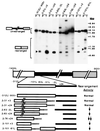
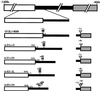
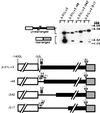
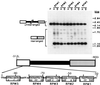
In the ciliate Tetrahymena thermophila, thousands of DNA segments of variable size are eliminated from the developing somatic macronucleus by specific DNA rearrangements. It is unclear whether rearrangement of the many different DNA elements occurs via a single mechanism or via multiple rearrangement systems. In this study, we characterized in vivo cis-acting sequences required for the rearrangement of the 1.1-kbp R deletion element. We found that rearrangement requires specific sequences flanking each side of the deletion element. The required sequences on the left side appear to span roughly a 70-bp region that is located at least 30 bp from the rearrangement boundary. When we moved the location of the left cis-acting sequences closer to the eliminated region, we observed a rightward shift of the rearrangement boundary such that the newly formed deletion junction retained its original distance from this flanking region. Likewise, when we moved the flanking region as much as 500 bp away from the deletion element, the rearrangement boundary shifted to remain in relative juxtaposition. Clusters of base substitutions made throughout this critical flanking region did not affect rearrangement efficiency or accuracy, which suggests a complex nature for this regulatory sequence. We also found that the right flanking region effectively replaced the essential sequences identified on the left side, and thus, the two flanking regions contain sequences of analogous function despite the lack of obvious sequence identity. These data taken together indicate that the R-element flanking regions contain sequences that position the rearrangement boundaries from a short distance away. Previously, a 10-bp polypurine tract flanking the M-deletion element was demonstrated to act from a distance to determine its rearrangement boundaries. No apparent sequence similarity exists between the M and R elements. The functional similarity between these different cis-acting sequences of the two elements is firm support for a common mechanism controlling Tetrahymena rearrangement.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
