Different mismatch repair deficiencies all have the same effects on somatic hypermutation: intact primary mechanism accompanied by secondary modifications
- PMID: 10429667
- PMCID: PMC2195558
- DOI: 10.1084/jem.190.1.21
Different mismatch repair deficiencies all have the same effects on somatic hypermutation: intact primary mechanism accompanied by secondary modifications
Abstract
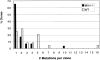
Somatic hypermutation of Ig genes is probably dependent on transcription of the target gene via a mutator factor associated with the RNA polymerase (Storb, U., E.L. Klotz, J. Hackett, Jr., K. Kage, G. Bozek, and T.E. Martin. 1998. J. Exp. Med. 188:689-698). It is also probable that some form of DNA repair is involved in the mutation process. It was shown that the nucleotide excision repair proteins were not required, nor were mismatch repair (MMR) proteins. However, certain changes in mutation patterns and frequency of point mutations were observed in Msh2 (MutS homologue) and Pms2 (MutL homologue) MMR-deficient mice (for review see Kim, N., and U. Storb. 1998. J. Exp. Med. 187:1729-1733). These data were obtained from endogenous immunoglobulin (Ig) genes and were presumably influenced by selection of B cells whose Ig genes had undergone certain mutations. In this study, we have analyzed somatic hypermutation in two MutL types of MMR deficiencies, Pms2 and Mlh1. The mutation target was a nonselectable Ig-kappa gene with an artificial insert in the V region. We found that both Pms2- and Mlh1-deficient mice can somatically hypermutate the Ig test gene at approximately twofold reduced frequencies. Furthermore, highly mutated sequences are almost absent. Together with the finding of genome instability in the germinal center B cells, these observations support the conclusion, previously reached for Msh2 mice, that MMR-deficient B cells undergoing somatic hypermutation have a short life span. Pms2- and Mlh-1-deficient mice also resemble Msh2-deficient mice with respect to preferential targeting of G and C nucleotides. Thus, it appears that the different MMR proteins do not have unique functions with respect to somatic hypermutation. Several intrinsic characteristics of somatic hypermutation remain unaltered in the MMR-deficient mice: a preference for targeting A over T, a strand bias, mutational hot spots, and hypermutability of the artificial insert are all seen in the unselectable Ig gene. This implies that the MMR proteins are not required for and most likely are not involved in the primary step of introducing the mutations. Instead, they are recruited to repair certain somatic point mutations, presumably soon after these are created.
Figures

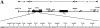
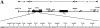


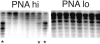



Similar articles
-
Mlh1 can function in antibody class switch recombination independently of Msh2.J Exp Med. 2003 May 19;197(10):1377-83. doi: 10.1084/jem.20022190. Epub 2003 May 12. J Exp Med. 2003. PMID: 12743174 Free PMC article.
-
Hypermutation in Ig V genes from mice deficient in the MLH1 mismatch repair protein.J Immunol. 1999 Mar 15;162(6):3121-4. J Immunol. 1999. PMID: 10092760
-
Role for mismatch repair proteins Msh2, Mlh1, and Pms2 in immunoglobulin class switching shown by sequence analysis of recombination junctions.J Exp Med. 2002 Feb 4;195(3):367-73. doi: 10.1084/jem.20011877. J Exp Med. 2002. PMID: 11828012 Free PMC article.
-
Cis-acting sequences that affect somatic hypermutation of Ig genes.Immunol Rev. 1998 Apr;162:153-60. doi: 10.1111/j.1600-065x.1998.tb01438.x. Immunol Rev. 1998. PMID: 9602361 Review.
-
Molecular basis of HNPCC: mutations of MMR genes.Hum Mutat. 1997;10(2):89-99. doi: 10.1002/(SICI)1098-1004(1997)10:2<89::AID-HUMU1>3.0.CO;2-H. Hum Mutat. 1997. PMID: 9259192 Review.
Cited by
-
DNA polymerase iota and related rad30-like enzymes.Philos Trans R Soc Lond B Biol Sci. 2001 Jan 29;356(1405):53-60. doi: 10.1098/rstb.2000.0748. Philos Trans R Soc Lond B Biol Sci. 2001. PMID: 11205331 Free PMC article. Review.
-
I. VH gene transcription creates stabilized secondary structures for coordinated mutagenesis during somatic hypermutation.Mol Immunol. 2008 Aug;45(13):3589-99. doi: 10.1016/j.molimm.2008.02.030. Epub 2008 Jun 27. Mol Immunol. 2008. PMID: 18585784 Free PMC article.
-
Mlh1 can function in antibody class switch recombination independently of Msh2.J Exp Med. 2003 May 19;197(10):1377-83. doi: 10.1084/jem.20022190. Epub 2003 May 12. J Exp Med. 2003. PMID: 12743174 Free PMC article.
-
The ATPase activity of MLH1 is required to orchestrate DNA double-strand breaks and end processing during class switch recombination.J Exp Med. 2012 Apr 9;209(4):671-8. doi: 10.1084/jem.20111531. Epub 2012 Mar 26. J Exp Med. 2012. PMID: 22451719 Free PMC article.
-
H2AX is required for recombination between immunoglobulin switch regions but not for intra-switch region recombination or somatic hypermutation.J Exp Med. 2003 Jun 16;197(12):1767-78. doi: 10.1084/jem.20030569. J Exp Med. 2003. PMID: 12810694 Free PMC article.
References
-
- Kelsoe G. The germinal center reaction. Immunol. Today. 1995;16:324–326. - PubMed
-
- Storb U. The molecular basis of somatic hypermutation of immunoglobulin genes. Curr. Opin. Immunol. 1996;8:206–214. - PubMed
-
- Storb U., Peters A., Klotz E., Kim N., Shen H.M., Kage K., Rogerson B. Somatic hypermutation of immunoglobulin genes is linked to transcription. Curr. Top. Microbiol. Immunol. 1998;229:11–19. - PubMed
-
- Doerner T., Brezinschek H.-P., Brezinschek R., Foster S., Domiati-Saad R., Lipsky P. Analysis of the frequency and pattern of somatic mutations within nonproductively rearranged human variable heavy chain genes. J. Immunol. 1997;158:2779–2789. - PubMed
-
- Smith D., Creadon G., Jena P., Portanova J., Kotzin B., Wysocki L. Di- and trinucleotide target preferences of somatic mutagenesis in normal and autoreactive B cells. J. Immunol. 1996;156:2642–2652. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

