Characterization and catalytic properties of the sterol 14alpha-demethylase from Mycobacterium tuberculosis
- PMID: 10430874
- PMCID: PMC17711
- DOI: 10.1073/pnas.96.16.8937
Characterization and catalytic properties of the sterol 14alpha-demethylase from Mycobacterium tuberculosis
Abstract
Sterol 14alpha-demethylase encoded by CYP51 is a mixed-function oxidase involved in sterol synthesis in eukaryotic organisms. Completion of the Mycobacterium tuberculosis genome project revealed that a protein having homology to mammalian 14alpha-demethylases might be present in this bacterium. Using genomic DNA from mycobacterial strain H(37)Rv, we have established unambiguously that the CYP51-like gene encodes a bacterial sterol 14alpha-demethylase. Expression of the M. tuberculosis CYP51 gene in Escherichia coli yields a P450, which, when purified to homogeneity, has the predicted molecular mass, ca. 50 kDa on SDS/PAGE, and binds both sterol substrates and azole inhibitors of P450 14alpha-demethylases. It catalyzes 14alpha-demethylation of lanosterol, 24, 25-dihydrolanosterol, and obtusifoliol to produce the 8,14-dienes stereoselectively as shown by GC/MS and (1)H NMR analysis. Both flavodoxin and ferredoxin redox systems are able to support this enzymatic activity. Structural requirements of a 14alpha-methyl group and Delta(8(9))-bond were established by comparing binding of pairs of sterol substrate that differed in a single molecular feature, e.g., cycloartenol paired with lanosterol. These substrate requirements are similar to those established for plant and animal P450 14alpha-demethylases. From the combination of results, the interrelationships of substrate functional groups within the active site show that oxidative portions of the sterol biosynthetic pathway are present in prokaryotes.
Figures

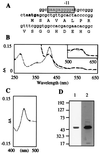



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

