A role for amontillado, the Drosophila homolog of the neuropeptide precursor processing protease PC2, in triggering hatching behavior
- PMID: 10436051
- PMCID: PMC6782853
- DOI: 10.1523/JNEUROSCI.19-16-06942.1999
A role for amontillado, the Drosophila homolog of the neuropeptide precursor processing protease PC2, in triggering hatching behavior
Abstract
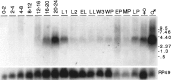
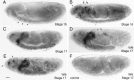
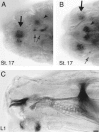
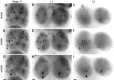
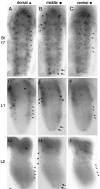
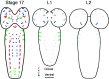
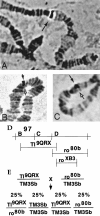
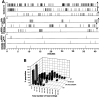
Accurate proteolytic processing of neuropeptide and peptide hormone precursors by members of the kexin/furin family of proteases is key to determining both the identities and activities of signaling peptides. Here we identify amontillado (amon), the Drosophila melanogaster homolog of the mammalian neuropeptide processing protease PC2, and show that in contrast to vertebrate PC2, amontillado expression undergoes extensive regulation in the nervous system during development. In situ hybridization reveals that expression of amontillado is restricted to the final stages of embryogenesis when it is found in anterior sensory structures and in only 168 cells in the brain and ventral nerve cord. After larvae hatch from their egg shells, the sensory structures and most cells in the CNS turn off or substantially reduce amontillado expression, suggesting that amontillado plays a specific role late in embryogenesis. Larvae lacking the chromosomal region containing amontillado show no gross anatomical defects and respond to touch. However, such larvae show a greatly reduced frequency of a hatching behavior of wild-type Drosophila in which larvae swing their heads, scraping through the eggshell with their mouth hooks. Ubiquitous expression of amontillado can restore near wild-type levels of this behavior, whereas expression of amontillado with an alanine substitution for the catalytic histidine cannot. These results suggest that amontillado expression is regulated as part of a programmed modulation of neural signaling that controls hatching behavior by producing specific neuropeptides in particular neurons at an appropriate developmental time.
Figures

Similar articles
-
The proprotein convertase encoded by amontillado (amon) is required in Drosophila corpora cardiaca endocrine cells producing the glucose regulatory hormone AKH.PLoS Genet. 2010 May 27;6(5):e1000967. doi: 10.1371/journal.pgen.1000967. PLoS Genet. 2010. PMID: 20523747 Free PMC article.
-
amontillado, the Drosophila homolog of the prohormone processing protease PC2, is required during embryogenesis and early larval development.Genetics. 2003 Jan;163(1):227-37. doi: 10.1093/genetics/163.1.227. Genetics. 2003. PMID: 12586710 Free PMC article.
-
Interaction of Drosophila melanogaster prohormone convertase 2 and 7B2. Insect cell-specific processing and secretion.J Biol Chem. 2000 Jun 9;275(23):17886-93. doi: 10.1074/jbc.M000032200. J Biol Chem. 2000. PMID: 10749852
-
Structure and function of eukaryotic proprotein processing enzymes of the subtilisin family of serine proteases.Crit Rev Oncog. 1993;4(2):115-36. Crit Rev Oncog. 1993. PMID: 8420571 Review.
-
The family of subtilisin/kexin like pro-protein and pro-hormone convertases: divergent or shared functions.Biochimie. 1994;76(3-4):197-209. doi: 10.1016/0300-9084(94)90147-3. Biochimie. 1994. PMID: 7819324 Review.
Cited by
-
Repression of liver colorectal metastasis by the serpin Spn4A a naturally occurring inhibitor of the constitutive secretory proprotein convertases.Oncotarget. 2014 Jun 30;5(12):4195-210. doi: 10.18632/oncotarget.1966. Oncotarget. 2014. PMID: 24961901 Free PMC article.
-
Multiple amidated neuropeptides are required for normal circadian locomotor rhythms in Drosophila.J Neurosci. 2001 Sep 1;21(17):6673-86. doi: 10.1523/JNEUROSCI.21-17-06673.2001. J Neurosci. 2001. PMID: 11517257 Free PMC article.
-
The EGL-3 proprotein convertase regulates mechanosensory responses of Caenorhabditis elegans.J Neurosci. 2001 Dec 1;21(23):9265-72. doi: 10.1523/JNEUROSCI.21-23-09265.2001. J Neurosci. 2001. PMID: 11717360 Free PMC article.
-
Genome-wide features of neuroendocrine regulation in Drosophila by the basic helix-loop-helix transcription factor DIMMED.Nucleic Acids Res. 2015 Feb 27;43(4):2199-215. doi: 10.1093/nar/gku1377. Epub 2015 Jan 29. Nucleic Acids Res. 2015. PMID: 25634895 Free PMC article.
-
The proprotein convertase encoded by amontillado (amon) is required in Drosophila corpora cardiaca endocrine cells producing the glucose regulatory hormone AKH.PLoS Genet. 2010 May 27;6(5):e1000967. doi: 10.1371/journal.pgen.1000967. PLoS Genet. 2010. PMID: 20523747 Free PMC article.
References
-
- Al-Atia GR, Fruscoloni P, Jacobs-Lorena M. Translational regulation of mRNAs for ribosomal proteins during early Drosophila development. Biochemistry. 1985;24:5798–5803. - PubMed
-
- Andrews PC, Brayton KA, Dixon JE. Post-translational proteolytic processing of precursors to regulatory peptides. Experientia [Suppl] 1989;56:192–209. - PubMed
-
- Ashburner M. Drosophila: a laboratory handbook. Cold Spring Harbor Laboratory; Cold Spring Harbor, New York: 1989.
-
- Bernheim SM, Mayeri E. Complex behavior induced by egg-laying hormone in Aplysia. J Comp Physiol [A] 1995;176:131–136. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
