Characterization of MdMADS2, a member of the SQUAMOSA subfamily of genes, in apple
- PMID: 10444080
- PMCID: PMC59356
- DOI: 10.1104/pp.120.4.969
Characterization of MdMADS2, a member of the SQUAMOSA subfamily of genes, in apple
Abstract
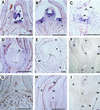
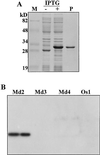
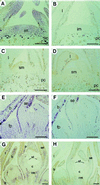
A MADS-box gene, MdMADS2, was isolated from the apple (Malus x domestica Borkh.) var Fuji and its developmental expression pattern was studied during flower development. MdMADS2 shares a high degree of amino acid sequence identity with the SQUAMOSA subfamily of genes. RNA blot analysis showed that MdMADS2 is transcribed through all stages of flower development, and its transcription was seen in the four floral organs. RNA in situ hybridization revealed that the MdMADS2 mRNA is expressed both in the inflorescence meristem and in the floral meristem. The MdMADS2 transcript was detected at all stages of flower development. Protein localization analysis showed that MdMADS2 protein was excluded from the stamen and carpel primordia, in which a considerable MdMADS2 mRNA signal was detected. This indicates that posttanscriptional regulation may be involved in the MdMADS2-mediated control of flower development. Transgenic tobacco expressing the MdMADS2 gene from the cauliflower mosaic virus 35S promoter showed early flowering and shorter bolts, but did not show any homeotic changes in the floral organs. These results suggest that MdMADS2 plays an important role during early stages of flower development.
Figures




Similar articles
-
Developmentally regulated expression of two MADS-box genes, MdMADS3 and MdMADS4, in the morphogenesis of flower buds and fruits in apple.Planta. 2000 Mar;210(4):519-28. doi: 10.1007/s004250050040. Planta. 2000. PMID: 10787044
-
Characterization of the potato MADS-box gene STMADS16 and expression analysis in tobacco transgenic plants.Plant Mol Biol. 2000 Feb;42(3):499-513. doi: 10.1023/a:1006397427894. Plant Mol Biol. 2000. PMID: 10798619
-
A TM3-like MADS-box gene from Eucalyptus expressed in both vegetative and reproductive tissues.Gene. 1999 Mar 4;228(1-2):155-60. doi: 10.1016/s0378-1119(98)00613-1. Gene. 1999. PMID: 10072768
-
Control of floral organ identity by homeotic MADS-box transcription factors.Results Probl Cell Differ. 1994;20:235-58. doi: 10.1007/978-3-540-48037-2_11. Results Probl Cell Differ. 1994. PMID: 7913550 Review. No abstract available.
-
Biochemical and molecular genetic aspects of floral scents.Plant Physiol. 2000 Mar;122(3):627-33. doi: 10.1104/pp.122.3.627. Plant Physiol. 2000. PMID: 10712525 Free PMC article. Review. No abstract available.
Cited by
-
The MADS box gene FBP2 is required for SEPALLATA function in petunia.Plant Cell. 2003 Apr;15(4):914-25. doi: 10.1105/tpc.010280. Plant Cell. 2003. PMID: 12671087 Free PMC article.
-
Overexpression of two PsnAP1 genes from Populus simonii × P. nigra causes early flowering in transgenic tobacco and Arabidopsis.PLoS One. 2014 Oct 31;9(10):e111725. doi: 10.1371/journal.pone.0111725. eCollection 2014. PLoS One. 2014. PMID: 25360739 Free PMC article.
-
Novel anther-specific myb genes from tobacco as putative regulators of phenylalanine ammonia-lyase expression.Plant Physiol. 2001 Aug;126(4):1738-53. doi: 10.1104/pp.126.4.1738. Plant Physiol. 2001. PMID: 11500571 Free PMC article.
-
A Global View of Transcriptome Dynamics During Male Floral Bud Development in Populus tomentosa.Sci Rep. 2018 Jan 15;8(1):722. doi: 10.1038/s41598-017-18084-5. Sci Rep. 2018. PMID: 29335419 Free PMC article.
-
The MADS and the Beauty: Genes Involved in the Development of Orchid Flowers.Curr Genomics. 2011 Aug;12(5):342-56. doi: 10.2174/138920211796429754. Curr Genomics. 2011. PMID: 22294877 Free PMC article.
References
-
- Amasino RM. Control of flowering time in plants. Curr Opin Genet Dev. 1996;6:480–487. - PubMed
-
- An G, Ebert PR, Mitra A, Ha S-B. Binary vectors. In: Gelvin SB, Schilperoort R, editors. Plant Molecular Biology Manual. Dordrecht, The Netherlands: Kluwer Academic Publishers; 1988. , section A3, pp 1–19.
-
- Angenent GC, Franken J, Busscher M, Weiss D, van Tunen AJ. Co-suppression of the petunia homeotic gene fbp2 affects the identity of the generative meristem. Plant J. 1994;5:33–44. - PubMed
-
- Bernier G. The control of floral evocation and morphogenesis. Annu Rev Plant Physiol Plant Mol Biol. 1988;39:175–219.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

