The crystal structure of anthranilate synthase from Sulfolobus solfataricus: functional implications
- PMID: 10449718
- PMCID: PMC22234
- DOI: 10.1073/pnas.96.17.9479
The crystal structure of anthranilate synthase from Sulfolobus solfataricus: functional implications
Abstract
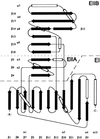
Anthranilate synthase catalyzes the synthesis of anthranilate from chorismate and glutamine and is feedback-inhibited by tryptophan. The enzyme of the hyperthermophile Sulfolobus solfataricus has been crystallized in the absence of physiological ligands, and its three-dimensional structure has been determined at 2.5-A resolution with x-ray crystallography. It is a heterotetramer of anthranilate synthase (TrpE) and glutamine amidotransferase (TrpG) subunits, in which two TrpG:TrpE protomers associate mainly via the TrpG subunits. The small TrpG subunit (195 residues) has the known "triad" glutamine amidotransferase fold. The large TrpE subunit (421 residues) has a novel fold. It displays a cleft between two domains, the tips of which contact the TrpG subunit across its active site. Clusters of catalytically essential residues are located inside the cleft, spatially separated from clustered residues involved in feedback inhibition. The structure suggests a model in which chorismate binding triggers a relative movement of the two domain tips of the TrpE subunit, activating the TrpG subunit and creating a channel for passage of ammonia toward the active site of the TrpE subunit. Tryptophan presumably blocks this rearrangement, thus stabilizing the inactive states of both subunits. The structure of the TrpE subunit is a likely prototype for the related enzymes 4-amino 4-deoxychorismate synthase and isochorismate synthase.
Figures



Similar articles
-
The structures of anthranilate synthase of Serratia marcescens crystallized in the presence of (i) its substrates, chorismate and glutamine, and a product, glutamate, and (ii) its end-product inhibitor, L-tryptophan.Proc Natl Acad Sci U S A. 2001 May 22;98(11):6021-6. doi: 10.1073/pnas.111150298. Proc Natl Acad Sci U S A. 2001. PMID: 11371633 Free PMC article.
-
Structure of the cooperative allosteric anthranilate synthase from Salmonella typhimurium.Nat Struct Biol. 2001 Mar;8(3):243-7. doi: 10.1038/84988. Nat Struct Biol. 2001. PMID: 11224570
-
Anthranilate synthase without an LLES motif from a hyperthermophilic archaeon is inhibited by tryptophan.Biochem Biophys Res Commun. 2001 Mar 9;281(4):858-65. doi: 10.1006/bbrc.2001.4428. Biochem Biophys Res Commun. 2001. PMID: 11237738
-
Conformational changes in ammonia-channeling glutamine amidotransferases.Curr Opin Struct Biol. 2007 Dec;17(6):653-64. doi: 10.1016/j.sbi.2007.09.003. Epub 2007 Oct 24. Curr Opin Struct Biol. 2007. PMID: 17951049 Review.
-
The amidotransferase family of enzymes: molecular machines for the production and delivery of ammonia.Biochemistry. 1999 Jun 22;38(25):7891-9. doi: 10.1021/bi990871p. Biochemistry. 1999. PMID: 10387030 Review.
Cited by
-
Allosteric pathways in imidazole glycerol phosphate synthase.Proc Natl Acad Sci U S A. 2012 May 29;109(22):E1428-36. doi: 10.1073/pnas.1120536109. Epub 2012 May 14. Proc Natl Acad Sci U S A. 2012. PMID: 22586084 Free PMC article.
-
Biosynthesis of the enediyne antitumor antibiotic C-1027 involves a new branching point in chorismate metabolism.Proc Natl Acad Sci U S A. 2008 Jan 15;105(2):494-9. doi: 10.1073/pnas.0708750105. Epub 2008 Jan 8. Proc Natl Acad Sci U S A. 2008. PMID: 18182490 Free PMC article.
-
Revitalizing antifolates through understanding mechanisms that govern susceptibility and resistance.Medchemcomm. 2019 May 8;10(6):880-895. doi: 10.1039/c9md00078j. eCollection 2019 Jun 1. Medchemcomm. 2019. PMID: 31303985 Free PMC article. Review.
-
Molecular basis for the allosteric activation mechanism of the heterodimeric imidazole glycerol phosphate synthase complex.Nat Commun. 2021 May 12;12(1):2748. doi: 10.1038/s41467-021-22968-6. Nat Commun. 2021. PMID: 33980881 Free PMC article.
-
Temperature-dependent function of the glutamine phosphoribosylpyrophosphate amidotransferase ammonia channel and coupling with glycinamide ribonucleotide synthetase in a hyperthermophile.J Bacteriol. 2000 Jul;182(13):3734-9. doi: 10.1128/JB.182.13.3734-3739.2000. J Bacteriol. 2000. PMID: 10850988 Free PMC article.
References
-
- Zalkin H. Adv Enzymol Relat Areas Mol Biol. 1993;66:203–309. - PubMed
-
- Zalkin H, Smith J L. Adv Enzymol Relat Areas Mol Biol. 1998;72:87–144. - PubMed
-
- Tesmer J J G, Klem T J, Deras M L, Davisson V J, Smith J L. Nat Struct Biol. 1996;3:74–86. - PubMed
-
- Caligiuri M G, Bauerle R. J Biol Chem. 1991;266:8328–8335. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

