The speed limit for protein folding measured by triplet-triplet energy transfer
- PMID: 10449738
- PMCID: PMC22254
- DOI: 10.1073/pnas.96.17.9597
The speed limit for protein folding measured by triplet-triplet energy transfer
Abstract
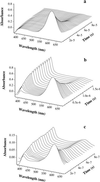
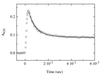
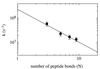
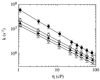
A direct measure of intramolecular chain diffusion is obtained by the determination of triplet-triplet energy-transfer rates between a donor and an acceptor chromophore attached at defined points on a polypeptide chain. Single exponential kinetics of contact formation are observed on the nanosecond time scale for polypeptides in which donor and acceptor are linked by repeating units of glycine and serine residues. The rates depend on the number of peptide bonds (N) separating donor and acceptor and show a maximum for the shortest peptides (N = 3) with a time constant (tau = 1/k) of 20 ns. This sets an upper limit for the speed of formation of the first side-chain contacts during protein folding.
Figures
References
-
- Kuwajima K. Proteins Struct Funct Genet. 1989;6:87–103. - PubMed
-
- Dobson C M. Curr Opin Struct Biol. 1992;2:6–12.
-
- Segel D, Bachmann A, Hofrichter J, Hodgson K, Doniach S, Kiefhaber T. J Mol Biol. 1999;288:489–500. - PubMed
-
- Jackson S E, Fersht A R. Biochemistry. 1991;30:10428–10435. - PubMed
-
- Alexander P, Orban J, Bryan P. Biochemistry. 1992;31:7243–7248. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

