Two major forms of DNA (cytosine-5) methyltransferase in human somatic tissues
- PMID: 10449766
- PMCID: PMC22282
- DOI: 10.1073/pnas.96.17.9751
Two major forms of DNA (cytosine-5) methyltransferase in human somatic tissues
Abstract
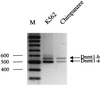
Thus far, only one major form of vertebrate DNA (cytosine-5) methyltransferase (CpG MTase, EC 2.1.1.37) has been identified, cloned, and extensively studied. This enzyme, dnmt1, has been hypothesized to be responsible for most of the maintenance as well as the de novo methylation activities occurring in the somatic cells of vertebrates. We now report the discovery of another abundant species of CpG MTase in various types of human cell lines and somatic tissues. Interestingly, the mRNA encoding this CpG MTase results from alternative splicing of the primary transcript from the Dnmt1 gene, which incorporates in-frame an additional 48 nt between exons 4 and 5. Furthermore, this 48-nt exon sequence is derived from the first, or the most upstream, copy of a set of seven different Alu repeats located in intron 4. The ratios of expression of this mRNA to the expression of the previously known, shorter Dnmt1 mRNA species, as estimated by semiquantitative reverse transcription-PCR analysis, range from two-thirds to three-sevenths. This alternative splicing scheme of the Dnmt1 transcript seems to be conserved in the higher primates. We suggest that the originally described and the recently discovered forms of CpG MTase be named dnmt1-a and dnmt1-b, respectively. The evolutionary and biological implications of this finding are discussed in relation to the cellular functions of the CpG residues and the CpG MTases.
Figures

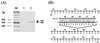

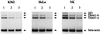



References
-
- Jost J P, Saluz H P, editors. DNA Methylation: Molecular Biology and Biological Significance. Basel: Birkhauser; 1993.
-
- Bestor T H, Verdine G L. Curr Opin Cell Biol. 1994;6:386–389. - PubMed
-
- Bestor T H. Nature (London) 1998;393:311–312. - PubMed
-
- Jones P A, Buckley J D. Adv Cancer Res. 1990;54:1–23. - PubMed
-
- Baylin S B. Science. 1997;277:1948–1949. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

