Fungal resistance to plant antibiotics as a mechanism of pathogenesis
- PMID: 10477313
- PMCID: PMC103751
- DOI: 10.1128/MMBR.63.3.708-724.1999
Fungal resistance to plant antibiotics as a mechanism of pathogenesis
Abstract
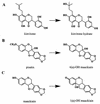
Many plants produce low-molecular-weight compounds which inhibit the growth of phytopathogenic fungi in vitro. These compounds may be preformed inhibitors that are present constitutively in healthy plants (also known as phytoanticipins), or they may be synthesized in response to pathogen attack (phytoalexins). Successful pathogens must be able to circumvent or overcome these antifungal defenses, and this review focuses on the significance of fungal resistance to plant antibiotics as a mechanism of pathogenesis. There is increasing evidence that resistance of fungal pathogens to plant antibiotics can be important for pathogenicity, at least for some fungus-plant interactions. This evidence has emerged largely from studies of fungal degradative enzymes and also from experiments in which plants with altered levels of antifungal secondary metabolites were generated. Whereas the emphasis to date has been on degradative mechanisms of resistance of phytopathogenic fungi to antifungal secondary metabolites, in the future we are likely to see a rapid expansion in our knowledge of alternative mechanisms of resistance. These may include membrane efflux systems of the kind associated with multidrug resistance and innate resistance due to insensitivity of the target site. The manipulation of plant biosynthetic pathways to give altered antibiotic profiles will also be valuable in telling us more about the significance of antifungal secondary metabolites for plant defense and clearly has great potential for enhancing disease resistance for commercial purposes.
Figures



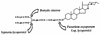
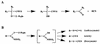

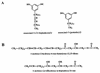
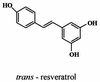
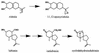

References
-
- Adrian M, Jeandet P, Veneau J, Weston L A, Bessis R. Biological activity of resveratrol, a stilbenic compound from grapevines, against Botrytis cinerea, the causal agent for gray mould. J Chem Ecol. 1997;23:1689–1702.
-
- Adrian M, Rajaei H, Jeandet P, Veneau J, Bessis R. Resveratrol oxidation in Botrytis cinerea conidia. Phytopathology. 1998;88:472–476. - PubMed
-
- Ardi R, Kobiler I, Jacoby B, Keen N T, Prusky D. Involvement of epicatechin biosynthesis in the activation of the mechanism of resistance of avocado fruits to Colletotrichum gloeosporioides. Physiol Mol Plant Pathol. 1998;53:269–285.
-
- Argandoña V H, Corcuera L J. Distribution of hydroxamic acids in Zea mays tissues. Phytochemistry. 1985;24:177–178.
-
- Argandoña V H, Zúniga G E, Corcuera L J. Distribution of gramine and hydroxamic acids in barley and wheat leaves. Phytochemistry. 1987;26:1917–1918.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

