DOT4 links silencing and cell growth in Saccharomyces cerevisiae
- PMID: 10490600
- PMCID: PMC84633
- DOI: 10.1128/MCB.19.10.6608
DOT4 links silencing and cell growth in Saccharomyces cerevisiae
Abstract
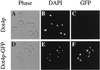
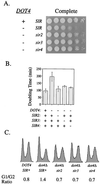
Transcriptional silencing in Saccharomyces cerevisiae occurs at specific loci and is mediated by a multiprotein complex that includes Rap1p and the Sir proteins. We studied the function of a recently identified gene, DOT4, that disrupts silencing when overexpressed. DOT4 encodes an ubiquitin processing protease (hydrolase) that is primarily located in the nucleus. By two-hybrid analysis, the amino-terminal third of Dot4p interacts with the silencing protein Sir4p. Cells lacking DOT4 exhibited reduced silencing and a corresponding decrease in the level of Sir4p. Together, these findings suggest that Dot4p regulates silencing by acting on Sir4p. In strains with several auxotrophic markers, loss of DOT4 ubiquitin hydrolase activity also results in a slow-growth defect. The defect can be partially suppressed by mutations in a subunit of the 26S proteasome, suggesting that Dot4p has the ability to prevent ubiquitin-mediated degradation. Furthermore, wild-type SIR2, SIR3, and SIR4 are required for full manifestation of the growth defect in a dot4 strain, indicating that the growth defect is caused in part by a silencing-related mechanism. We propose that Dot4p helps to restrict the location of silencing proteins to a limited set of genomic loci.
Figures






References
-
- Adams A, Gottschling D E, Kaiser C A, Stearns T. Methods in yeast genetics. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1997.
-
- Aparicio O M, Billington B L, Gottschling D E. Modifiers of position effect are shared between telomeric and silent mating-type loci in S. cerevisiae. Cell. 1991;66:1279–1287. - PubMed
-
- Aparicio O M, Gottschling D E. Overcoming telomeric silencing: a trans-activator competes to establish gene expression in a cell cycle-dependent way. Genes Dev. 1994;8:1133–1146. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D O, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: John Wiley & Sons; 1998.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
