The CCR4 and CAF1 proteins of the CCR4-NOT complex are physically and functionally separated from NOT2, NOT4, and NOT5
- PMID: 10490603
- PMCID: PMC84645
- DOI: 10.1128/MCB.19.10.6642
The CCR4 and CAF1 proteins of the CCR4-NOT complex are physically and functionally separated from NOT2, NOT4, and NOT5
Abstract
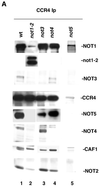
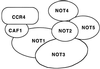
The CCR4-NOT complex (1 mDa in size), consisting of the proteins CCR4, CAF1, and NOT1 to NOT5, regulates gene expression both positively and negatively and is distinct from other large transcriptional complexes in Saccharomyces cerevisiae such as SNF/SWI, TFIID, SAGA, and RNA polymerase II holoenzyme. The physical and genetic interactions between the components of the CCR4-NOT complex were investigated in order to gain insight into how this complex affects the expression of diverse genes and processes. The CAF1 protein was found to be absolutely required for CCR4 association with the NOT proteins, and CCR4 and CAF1, in turn, physically interacted with NOT1 through its central amino acid region from positions 667 to 1152. The NOT3, NOT4, and NOT5 proteins had no significant effect on the association of CCR4, CAF1, and NOT1 with each other. In contrast, the NOT2, NOT4, and NOT5 interacted with the C-terminal region (residues 1490 to 2108) of NOT1 in which NOT2 and NOT5 physically associated in the absence of CAF1, NOT3, and NOT4. These and other data indicate that the physical ordering of these proteins in the complex is CCR4-CAF1-NOT1-(NOT2, NOT5), with NOT4 and NOT3 more peripheral to NOT2 and NOT5. The physical separation of CCR4 and CAF1 from other components of the CCR4-NOT complex correlated with genetic analysis indicating partially separate functions for these two groups of proteins. ccr4 or caf1 deletion suppressed the increased 3-aminotriazole resistance phenotype conferred by not mutations, resulted in opposite effects on gene expression as compared to several not mutations, and resulted in a number of synthetic phenotypes in combination with not mutations. These results define the CCR4-NOT complex as consisting of at least two physically and functionally separated groups of proteins.
Figures







References
-
- Carlson M. Genetics of transcriptional regulation in yeast: connections to the RNA polymerase II CTD. Annu Rev Cell Dev Biol. 1997;13:1–23. - PubMed
-
- Chen, J. Personal communication.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
