Functional characterization of rpn3 uncovers a distinct 19S proteasomal subunit requirement for ubiquitin-dependent proteolysis of cell cycle regulatory proteins in budding yeast
- PMID: 10490625
- PMCID: PMC84683
- DOI: 10.1128/MCB.19.10.6872
Functional characterization of rpn3 uncovers a distinct 19S proteasomal subunit requirement for ubiquitin-dependent proteolysis of cell cycle regulatory proteins in budding yeast
Abstract
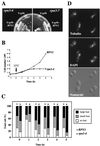
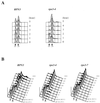
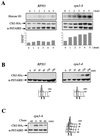
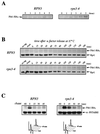
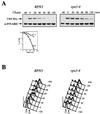
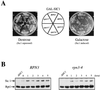
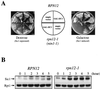
By selectively eliminating ubiquitin-conjugated proteins, the 26S proteasome plays a pivotal role in a large variety of cellular regulatory processes, particularly in the control of cell cycle transitions. Access of ubiquitinated substrates to the inner catalytic chamber within the 20S core particle is mediated by the 19S regulatory particle (RP), whose subunit composition in budding yeast has been recently elucidated. In this study, we have investigated the cell cycle defects resulting from conditional inactivation of one of these RP components, the essential non-ATPase Rpn3/Sun2 subunit. Using temperature-sensitive mutant alleles, we show that rpn3 mutations do not prevent the G(1)/S transition but cause a metaphase arrest, indicating that the essential Rpn3 function is limiting for mitosis. rpn3 mutants appear severely compromised in the ubiquitin-dependent proteolysis of several physiologically important proteasome substrates. Thus, RPN3 function is required for the degradation of the G(1)-phase cyclin Cln2 targeted by SCF; the S-phase cyclin Clb5, whose ubiquitination is likely to involve a combination of E3 (ubiquitin protein ligase) enzymes; and anaphase-promoting complex targets, such as the B-type cyclin Clb2 and the anaphase inhibitor Pds1. Our results indicate that the Pds1 degradation defect of the rpn3 mutants most likely accounts for the metaphase arrest phenotype observed. Surprisingly, but consistent with the lack of a G(1) arrest phenotype in thermosensitive rpn3 strains, the Cdk inhibitor Sic1 exhibits a short half-life regardless of the RPN3 genotype. In striking contrast, Sic1 turnover is severely impaired by a temperature-sensitive mutation in RPN12/NIN1, encoding another essential RP subunit. While other interpretations are possible, these data strongly argue for the requirement of distinct RP subunits for efficient proteolysis of specific cell cycle regulators. The potential implications of these data are discussed in the context of possible Rpn3 function in multiubiquitin-protein conjugate recognition by the 19S proteasomal regulatory particle.
Figures





References
-
- Amon A, Irniger S, Nasmyth K. Closing the cell cycle circle: G2 cyclin proteolysis initiated at mitosis persists until activation of G1 cyclins in the next cell cycle. Cell. 1994;77:1037–1050. - PubMed
-
- Bai C, Sen P, Hofmann K, Ma L, Goebl M, Harper J W, Elledge S J. SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell. 1996;86:263–274. - PubMed
-
- Bailly, E. Unpublished results.
-
- Barral Y, Jentsch S, Mann C. G1 cyclin turnover and nutrient uptake are controlled by a common pathway in yeast. Genes Dev. 1995;9:399–409. - PubMed
-
- Baumeister W, Walz J, Zhul F, Seemuller E. The proteasome: paradigm of a self-compartmentalizing protease. Cell. 1998;92:367–380. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
