Concerted regulation of wild-type p53 nuclear accumulation and activation by S100B and calcium-dependent protein kinase C
- PMID: 10490652
- PMCID: PMC84710
- DOI: 10.1128/MCB.19.10.7168
Concerted regulation of wild-type p53 nuclear accumulation and activation by S100B and calcium-dependent protein kinase C
Abstract
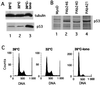
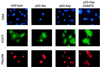
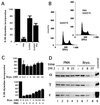
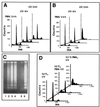
The calcium ionophore ionomycin cooperates with the S100B protein to rescue a p53-dependent G(1) checkpoint control in S100B-expressing mouse embryo fibroblasts and rat embryo fibroblasts (REF cells) which express the temperature-sensitive p53Val135 mutant (C. Scotto, J. C. Deloulme, D. Rousseau, E. Chambaz, and J. Baudier, Mol. Cell. Biol. 18:4272-4281, 1998). We investigated in this study the contributions of S100B and calcium-dependent PKC (cPKC) signalling pathways to the activation of wild-type p53. We first confirmed that S100B expression in mouse embryo fibroblasts enhanced specific nuclear accumulation of wild-type p53. We next demonstrated that wild-type p53 nuclear translocation and accumulation is dependent on cPKC activity. Mutation of the five putative cPKC phosphorylation sites on murine p53 into alanine or aspartic residues had no significant effect on p53 nuclear localization, suggesting that the cPKC effect on p53 nuclear translocation is indirect. A concerted regulation by S100B and cPKC of wild-type p53 nuclear translocation and activation was confirmed with REF cells expressing S100B (S100B-REF cells) overexpressing the temperature-sensitive p53Val135 mutant. Stimulation of S100B-REF cells with the PKC activator phorbol ester phorbol myristate acetate (PMA) promoted specific nuclear translocation of the wild-type p53Val135 species in cells positioned in early G(1) phase of the cell cycle. PMA also substituted for ionomycin in the mediating of p53-dependent G(1) arrest at the nonpermissive temperature (37.5 degrees C). PMA-dependent growth arrest was linked to the cell apoptosis response to UV irradiation. In contrast, growth arrest mediated by a temperature shift to 32 degrees C protected S100B-REF cells from apoptosis. Our results suggest a model in which calcium signalling, linked with cPKC activation, cooperates with S100B to promote wild-type p53 nuclear translocation in early G(1) phase and activation of a p53-dependent G(1) checkpoint control.
Figures







References
-
- Bates S, Vousden K. p53 in signalling checkpoint arrest or apoptosis. Curr Opin Genet Dev. 1996;6:12–19. - PubMed
-
- Baudier J, Glasser N, Gérard D. Ions binding to S100 protein. Calcium and zinc-binding properties of bovine brain S100αα, S100a (αβ), and S100B (ββ) protein: Zn2+ regulates Ca2+ binding on S100B protein. J Biol Chem. 1986;261:8192–8203. - PubMed
-
- Baudier J, Bergeret E, Bertacchi N, Weintraub H, Gagnon J, Garin J. Interactions of myogenic bHLH transcription factors with calcium-binding calmodulin and S100a (αα) proteins. Biochemistry. 1995;34:7834–7846. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
