Starvation induces vacuolar targeting and degradation of the tryptophan permease in yeast
- PMID: 10491387
- PMCID: PMC2156124
- DOI: 10.1083/jcb.146.6.1227
Starvation induces vacuolar targeting and degradation of the tryptophan permease in yeast
Abstract
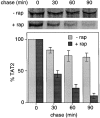
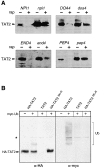
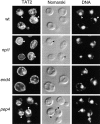
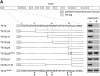
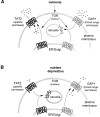
In Saccharomyces cerevisiae, amino acid permeases are divided into two classes. One class, represented by the general amino acid permease GAP1, contains permeases regulated in response to the nitrogen source. The other class, including the high affinity tryptophan permease, TAT2, consists of the so-called constitutive permeases. We show that TAT2 is regulated at the level of protein stability. In exponentially growing cells, TAT2 is in the plasma membrane and also accumulates in internal compartments of the secretory pathway. Upon nutrient deprivation or rapamycin treatment, TAT2 is transported to and degraded in the vacuole. The ubiquitination machinery and lysine residues within the NH(2)-terminal 31 amino acids of TAT2 mediate ubiquitination and degradation of the permease. Starvation-induced degradation of internal TAT2 is blocked in sec18, sec23, pep12, and vps27 mutants, but not in sec4, end4, and apg1 mutants, suggesting that, upon nutrient limitation, internal TAT2 is diverted from the late secretory pathway to the vacuolar pathway. Furthermore, our results suggest that TAT2 stability and sorting are controlled by the TOR signaling pathway, and regulated inversely to that of GAP1.
Figures
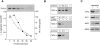



References
-
- Ausubel F.M., Brent R., Kingston R.E., Moore D.D., Seidman J.G., Smith J.A., Struhl K. Current Protocols in Molecular Biology. John Wiley & Sons, Inc; New York: 1998.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

