The ripX locus of Bacillus subtilis encodes a site-specific recombinase involved in proper chromosome partitioning
- PMID: 10498718
- PMCID: PMC103633
- DOI: 10.1128/JB.181.19.6053-6062.1999
The ripX locus of Bacillus subtilis encodes a site-specific recombinase involved in proper chromosome partitioning
Abstract
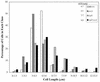
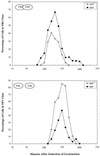
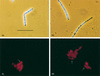
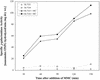
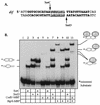
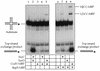
The Bacillus subtilis ripX gene encodes a protein that has 37 and 44% identity with the XerC and XerD site-specific recombinases of Escherichia coli. XerC and XerD are hypothesized to act in concert at the dif site to resolve dimeric chromosomes formed by recombination during replication. Cultures of ripX mutants contained a subpopulation of unequal-size cells held together in long chains. The chains included anucleate cells and cells with aberrantly dense or diffuse nucleoids, indicating a chromosome partitioning failure. This result is consistent with RipX having a role in the resolution of chromosome dimers in B. subtilis. Spores contain a single uninitiated chromosome, and analysis of germinated, outgrowing spores showed that the placement of FtsZ rings and septa is affected in ripX strains by the first division after the initiation of germination. The introduction of a recA mutation into ripX strains resulted in only slight modifications of the ripX phenotype, suggesting that chromosome dimers can form in a RecA-independent manner in B. subtilis. In addition to RipX, the CodV protein of B. subtilis shows extensive similarity to XerC and XerD. The RipX and CodV proteins were shown to bind in vitro to DNA containing the E. coli dif site. Together they functioned efficiently in vitro to catalyze site-specific cleavage of an artificial Holliday junction containing a dif site. Inactivation of codV alone did not cause a discernible change in phenotype, and it is speculated that RipX can substitute for CodV in vivo.
Figures

Similar articles
-
Identification and characterization of the dif Site from Bacillus subtilis.J Bacteriol. 2001 Feb;183(3):1058-68. doi: 10.1128/JB.183.3.1058-1068.2001. J Bacteriol. 2001. PMID: 11208805 Free PMC article.
-
Effects of replication termination mutants on chromosome partitioning in Bacillus subtilis.Proc Natl Acad Sci U S A. 2001 Jan 2;98(1):212-7. doi: 10.1073/pnas.98.1.212. Proc Natl Acad Sci U S A. 2001. PMID: 11134515 Free PMC article.
-
Sequential strand exchange by XerC and XerD during site-specific recombination at dif.J Biol Chem. 2000 Apr 7;275(14):9930-6. doi: 10.1074/jbc.275.14.9930. J Biol Chem. 2000. PMID: 10744667
-
A tale of two genomes: resolution of dimeric chromosomes in Escherichia coli and Bacillus subtilis.Res Microbiol. 2000 Sep;151(7):503-11. doi: 10.1016/s0923-2508(00)00220-5. Res Microbiol. 2000. PMID: 11037128 Review.
-
Circles: the replication-recombination-chromosome segregation connection.Proc Natl Acad Sci U S A. 2001 Jul 17;98(15):8189-95. doi: 10.1073/pnas.111008998. Proc Natl Acad Sci U S A. 2001. PMID: 11459952 Free PMC article. Review.
Cited by
-
Single-Molecule Tracking of DNA Translocases in Bacillus subtilis Reveals Strikingly Different Dynamics of SftA, SpoIIIE, and FtsA.Appl Environ Microbiol. 2018 Apr 2;84(8):e02610-17. doi: 10.1128/AEM.02610-17. Print 2018 Apr 15. Appl Environ Microbiol. 2018. PMID: 29439991 Free PMC article.
-
Effects of Phenotypic Variation on Biological Properties of Endophytic Bacteria Bacillus mojavensis PS17.Biology (Basel). 2022 Sep 2;11(9):1305. doi: 10.3390/biology11091305. Biology (Basel). 2022. PMID: 36138785 Free PMC article.
-
Comparison of responses to double-strand breaks between Escherichia coli and Bacillus subtilis reveals different requirements for SOS induction.J Bacteriol. 2009 Feb;191(4):1152-61. doi: 10.1128/JB.01292-08. Epub 2008 Dec 5. J Bacteriol. 2009. PMID: 19060143 Free PMC article.
-
XerD unloads bacterial SMC complexes at the replication terminus.Mol Cell. 2021 Feb 18;81(4):756-766.e8. doi: 10.1016/j.molcel.2020.12.027. Epub 2021 Jan 19. Mol Cell. 2021. PMID: 33472056 Free PMC article.
-
Tn917 targets the region where DNA replication terminates in Bacillus subtilis, highlighting a difference in chromosome processing in the firmicutes.J Bacteriol. 2009 Dec;191(24):7623-7. doi: 10.1128/JB.01023-09. Epub 2009 Oct 9. J Bacteriol. 2009. PMID: 19820088 Free PMC article.
References
-
- Blakely G, Colloms S D, May G, Burke M, Sherratt D J. Escherichia coli XerC recombinase is required for chromosomal segregation at cell division. New Biol. 1991;3:789–798. - PubMed
-
- Blakely G, May G, McCulloch R, Arciszewska L K, Burke M, Lovett S T, Sherratt D J. Two related recombinases are required for site-specific recombination at dif and cer in E. coli K12. Cell. 1993;75:351–361. - PubMed
-
- Blakely G, Sherratt D. Determinants of selectivity in Xer site-specific recombination. Genes Dev. 1996;10:762–773. - PubMed
-
- Blakely G W, Davidson A O, Sherratt D J. Binding and cleavage of nicked substrates by site-specific recombinases XerC and XerD. J Mol Biol. 1997;265:30–39. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

