Mutational analysis of the RecJ exonuclease of Escherichia coli: identification of phosphoesterase motifs
- PMID: 10498723
- PMCID: PMC103638
- DOI: 10.1128/JB.181.19.6098-6102.1999
Mutational analysis of the RecJ exonuclease of Escherichia coli: identification of phosphoesterase motifs
Abstract
The recJ gene, identified in Escherichia coli, encodes a Mg(+2)-dependent 5'-to-3' exonuclease with high specificity for single-strand DNA. Genetic and biochemical experiments implicate RecJ exonuclease in homologous recombination, base excision, and methyl-directed mismatch repair. Genes encoding proteins with strong similarities to RecJ have been found in every eubacterial genome sequenced to date, with the exception of Mycoplasma and Mycobacterium tuberculosis. Multiple genes encoding proteins similar to RecJ are found in some eubacteria, including Bacillus and Helicobacter, and in the archaea. Among this divergent set of sequences, seven conserved motifs emerge. We demonstrate here that amino acids within six of these motifs are essential for both the biochemical and genetic functions of E. coli RecJ. These motifs may define interactions with Mg(2+) ions or substrate DNA. A large family of proteins more distantly related to RecJ is present in archaea, eubacteria, and eukaryotes, including a hypothetical protein in the MgPa adhesin operon of Mycoplasma, a domain of putative polyA polymerases in Synechocystis and Aquifex, PRUNE of Drosophila, and an exopolyphosphatase (PPX1) of Saccharomyces cereviseae. Because these six RecJ motifs are shared between exonucleases and exopolyphosphatases, they may constitute an ancient phosphoesterase domain now found in all kingdoms of life.
Figures

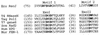
References
-
- Aravind L, Koonin E V. A novel family of predicted phosphoesterase includes Drosophila prune protein and bacterial RecJ exonuclease. Trends Biochem Sci. 1998;23:17–19. - PubMed
-
- Ausubel F A, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Short protocols in molecular biology. New York, N.Y: Greene Publishing and Wiley-Interscience; 1989.
-
- Bradford M M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

