Mapping of a hypovirus p29 protease symptom determinant domain with sequence similarity to potyvirus HC-Pro protease
- PMID: 10516056
- PMCID: PMC112982
- DOI: 10.1128/JVI.73.11.9478-9484.1999
Mapping of a hypovirus p29 protease symptom determinant domain with sequence similarity to potyvirus HC-Pro protease
Abstract
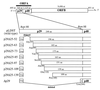
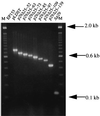
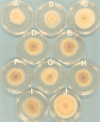
Hypovirus infection of the chestnut blight fungus Cryphonectria parasitica results in a spectrum of phenotypic changes that can include alterations in colony morphology and significant reductions in pigmentation, asexual sporulation, and virulence (hypovirulence). Deletion of 88% [Phe(25) to Pro(243)] of the virus-encoded papain-like protease, p29, in the context of an infectious cDNA clone of the prototypic hypovirus CHV1-EP713 (recombinant virus Deltap29) partially relieved virus-mediated suppression of pigmentation and sporulation without altering the level of hypovirulence. We now report mapping of the p29 symptom determinant domain to a region extending from Phe(25) through Gln(73) by a gain-of-function analysis following progressive repair of the Deltap29 deletion mutant. This domain was previously shown to share sequence similarity [including conserved cysteine residues Cys(38), Cys(48), Cys(70), and Cys(72)] with the N-terminal portion of the potyvirus-encoded helper component-proteinase (HC-Pro), a multifunctional protein implicated in aphid-mediated transmission, genome amplification, polyprotein processing, long-distance movement, and suppression of posttranscriptional silencing. Substitution of a glycine residue for either Cys(38) or Cys(48) resulted in no qualitative or quantitative changes in virus-mediated symptoms. Unexpectedly, mutation of Cys(70) resulted in a very severe phenotype that included significantly reduced mycelial growth and profoundly altered colony morphology. In contrast, substitution for Cys(72) resulted in a less severe symptom phenotype approaching that observed for Deltap29. The finding that p29-mediated symptom expression is influenced by two cysteine residues that are conserved in the potyvirus-encoded HC-Pro raises the possibility that these related viral-papain-like proteases function in their respective fungal and plant hosts by impacting ancestrally related regulatory pathways.
Figures

Similar articles
-
Hypovirus papain-like protease p29 functions in trans to enhance viral double-stranded RNA accumulation and vertical transmission.J Virol. 2003 Nov;77(21):11697-707. doi: 10.1128/jvi.77.21.11697-11707.2003. J Virol. 2003. PMID: 14557655 Free PMC article.
-
Synergism between a mycoreovirus and a hypovirus mediated by the papain-like protease p29 of the prototypic hypovirus CHV1-EP713.J Gen Virol. 2006 Dec;87(Pt 12):3703-3714. doi: 10.1099/vir.0.82213-0. J Gen Virol. 2006. PMID: 17098988
-
Characterization of the Papain-Like Protease p29 of the Hypovirus CHV1-CN280 in Its Natural Host Fungus Cryphonectria parasitica and Nonhost Fungus Magnaporthe oryzae.Phytopathology. 2019 May;109(5):736-747. doi: 10.1094/PHYTO-08-18-0318-R. Epub 2019 Apr 5. Phytopathology. 2019. PMID: 30592694
-
Expanding Repertoire of Plant Positive-Strand RNA Virus Proteases.Viruses. 2019 Jan 15;11(1):66. doi: 10.3390/v11010066. Viruses. 2019. PMID: 30650571 Free PMC article. Review.
-
3C-like proteases at the interface of plant-virus-vector interactions: Focus on potyvirid NIa proteases and secovirid proteases.Virology. 2025 Jan;602:110299. doi: 10.1016/j.virol.2024.110299. Epub 2024 Nov 14. Virology. 2025. PMID: 39579507 Review.
Cited by
-
Variations in hypovirus interactions with the fungal-host RNA-silencing antiviral-defense response.J Virol. 2012 Dec;86(23):12933-9. doi: 10.1128/JVI.00961-12. Epub 2012 Sep 19. J Virol. 2012. PMID: 22993160 Free PMC article.
-
Transfection of Sclerotinia sclerotiorum with in vitro transcripts of a naturally occurring interspecific recombinant of Sclerotinia sclerotiorum hypovirus 2 significantly reduces virulence of the fungus.J Virol. 2015 May;89(9):5060-71. doi: 10.1128/JVI.03199-14. Epub 2015 Feb 18. J Virol. 2015. PMID: 25694604 Free PMC article.
-
The HCPro from the Potyviridae family: an enviable multitasking Helper Component that every virus would like to have.Mol Plant Pathol. 2018 Mar;19(3):744-763. doi: 10.1111/mpp.12553. Epub 2017 May 26. Mol Plant Pathol. 2018. PMID: 28371183 Free PMC article. Review.
-
RNA interference pathways in fungi: mechanisms and functions.Annu Rev Microbiol. 2012;66:305-23. doi: 10.1146/annurev-micro-092611-150138. Epub 2012 Jun 28. Annu Rev Microbiol. 2012. PMID: 22746336 Free PMC article. Review.
-
Mycoreovirus genome alterations: similarities to and differences from rearrangements reported for other reoviruses.Front Microbiol. 2012 Jun 1;3:186. doi: 10.3389/fmicb.2012.00186. eCollection 2012. Front Microbiol. 2012. PMID: 22675320 Free PMC article.
References
-
- Anagnostakis S L. Biological control of chestnut blight. Science. 1982;215:466–471. - PubMed
-
- Atreya C D, Atreya P L, Thornbury D W, Pirone T P. Site-directed mutations in the potyvirus HC-PRO gene affect helper component activity, virus accumulation, and symptom expression in infected tobacco plants. Virology. 1992;191:106–111. - PubMed
-
- Chen B, Chen C-H, Bowman B H, Nuss D L. Phenotypic changes associated with wild-type and mutant hypovirus RNA transfection of plant pathogenic fungi phylogenetically related to Cryphonectria parasitica. Phytopathology. 1996;86:301–310.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

