A resource of mapped human bacterial artificial chromosome clones
- PMID: 10523527
- PMCID: PMC310825
- DOI: 10.1101/gr.9.10.989
A resource of mapped human bacterial artificial chromosome clones
Abstract
To date, despite the increasing number of genomic tools, there is no repository of ordered human BAC clones that covers entire chromosomes. This project presents a resource of mapped large DNA fragments that span eight human chromosomes at approximately 1-Mb resolution. These DNA fragments are bacterial artificial chromosome (BAC) clones anchored to sequence tagged site (STS) markers. This clone collection, which currently contains 759 mapped clones, is useful in a wide range of applications from microarray-based gene mapping to identification of chromosomal mutations. In addition to the clones themselves, we describe a database, GenMapDB (http://genomics.med.upenn.edu/genmapdb), that contains information about each clone in our collection.
Figures
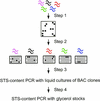
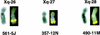

References
-
- Cheung VG, Nelson SF. Genomic mismatch scanning identifies human genomic DNA shared identical-by-descent. Genomics. 1998;47:1–7. - PubMed
-
- Cheung VG, Gregg JP, Gogolin-Ewens KJ, Bandong J, Stanley CA, Baker L, Higgins MJ, Nowak NJ, Shows TB, Ewens WJ, et al. Linkage disequilibrium mapping without genotyping. Nat Genet. 1998;18:225–230. - PubMed
-
- Collins FS, Patrinos A, Jordan E, Chakravarti A, Gesteland R, Walters L. New goals for the U.S. Human Genome Project: 1998–2003. Science. 1998;282:682–689. - PubMed
-
- Deloukas P, Schuler GD, Gyapay G, Beasley EM, Soderlund C, Rodriguez-Tome P, Hui L, Matise TC, McKusick KB, Beckmann JS, et al. A physical map of 30,000 human genes. Science. 1998;282:744–745. - PubMed
-
- Gyapay G, Schmitt K, Fizames C, Jones H, Vega-Czarny N, Spillett D, Muselet D, Prud'Homme JF, Dib C, Auffray C, et al. A radiation hybrid map of the human genome. Hum Mol Genet. 1996;5:339–346. - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
