Reciprocal secretion of proteins by the bacterial type III machines of plant and animal pathogens suggests universal recognition of mRNA targeting signals
- PMID: 10536009
- PMCID: PMC23122
- DOI: 10.1073/pnas.96.22.12839
Reciprocal secretion of proteins by the bacterial type III machines of plant and animal pathogens suggests universal recognition of mRNA targeting signals
Abstract
Bacterial pathogens of both animals and plants use type III secretion machines to inject virulence proteins into host cells. Although many components of the secretion machinery are conserved among different bacterial species, the substrates for their type III pathways are not. The Yersinia type III machinery recognizes some secretion substrates via a signal that is encoded within the first 15 codons of yop mRNA. These signals can be altered by frameshift mutations without affecting secretion of the encoded polypeptides, suggesting a mechanism whereby translation of yop mRNA is coupled to the translocation of newly synthesized polypeptide. We report that the type III machinery of Erwinia chrysanthemi cloned in Escherichia coli recognizes the secretion signals of yopE and yopQ. Pseudomonas syringae AvrB and AvrPto, two proteins exported by the recombinant Erwinia machine, can also be secreted by the Yersinia type III pathway. Mapping AvrPto sequences sufficient for the secretion of reporter fusions in Yersinia revealed the presence of an mRNA secretion signal. We propose that 11 conserved components of type III secretion machines may recognize signals that couple mRNA translation to polypeptide secretion.
Figures


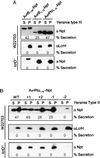

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

