Genetic ablation of root cap cells in Arabidopsis
- PMID: 10536027
- PMCID: PMC23173
- DOI: 10.1073/pnas.96.22.12941
Genetic ablation of root cap cells in Arabidopsis
Abstract
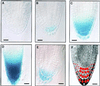
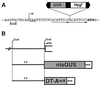
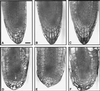
The root cap is increasingly appreciated as a complex and dynamic plant organ. Root caps sense and transmit environmental signals, synthesize and secrete small molecules and macromolecules, and in some species shed metabolically active cells. However, it is not known whether root caps are essential for normal shoot and root development. We report the identification of a root cap-specific promoter and describe its use to genetically ablate root caps by directing root cap-specific expression of a diphtheria toxin A-chain gene. Transgenic toxin-expressing plants are viable and have normal aerial parts but agravitropic roots, implying loss of root cap function. Several cell layers are missing from the transgenic root caps, and the remaining cells are abnormal. Although the radial organization of the roots is normal in toxin-expressing plants, the root tips have fewer cytoplasmically dense cells than do wild-type root tips, suggesting that root meristematic activity is lower in transgenic than in wild-type plants. The roots of transgenic plants have more lateral roots and these are, in turn, more highly branched than those of wild-type plants. Thus, root cap ablation alters root architecture both by inhibiting root meristematic activity and by stimulating lateral root initiation. These observations imply that the root caps contain essential components of the signaling system that determines root architecture.
Figures




References
-
- Gregory P J, Lake J V, Rose D A. Root Development and Function. Cambridge, U.K.: Cambridge Univ. Press; 1987.
-
- Bengough A G, McKenzie B M. J Exp Bot. 1997;48:885–893.
-
- Rogers H T, Pearson R W, Pierre W H. Soil Sci. 1942;54:353–356.
-
- Hawes M C, Brigham L A. Adv Plant Pathol. 1992;8:119–148.
-
- Rougier M. In: Plant Carbohydrates II. Tanner F A, Loewes F A, editors. 13B. Berlin: Springer; 1981. pp. 542–574.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

