Molecular evolution of the pathogenicity island of enterotoxigenic Bacteroides fragilis strains
- PMID: 10542162
- PMCID: PMC94125
- DOI: 10.1128/JB.181.21.6623-6633.1999
Molecular evolution of the pathogenicity island of enterotoxigenic Bacteroides fragilis strains
Abstract
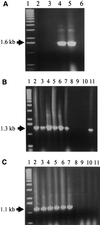
Enterotoxigenic Bacteroides fragilis (ETBF) strains, which produce a 20-kDa zinc metalloprotease toxin (BFT), have been associated with diarrheal disease in animals and young children. Studying a collection of ETBF and nontoxigenic B. fragilis (NTBF) strains, we found that bft and a second metalloprotease gene (mpII) are contained in an approximately 6-kb pathogenicity island (termed B. fragilis pathogenicity island or BfPAI) which is present exclusively in all 113 ETBF strains tested (pattern I). Of 191 NTBF strains, 100 (52%) lack both the BfPAI and at least a 12-kb region flanking BfPAI (pattern II), and 82 of 191 NTBF strains (43%) lack the BfPAI but contain the flanking region (pattern III). The nucleotide sequence flanking the left end of the BfPAI revealed a region with the same organization as the mobilization region of the 5-nitroimidazole resistance plasmid pIP417 and the clindamycin resistance plasmid pBFTM10, that is, two mobilization genes (bfmA and bfmB) organized in one operon and a putative origin of transfer (oriT) located in a small, compact region. The region flanking the right end of the BfPAI contains a gene (bfmC) whose predicted protein shares significant identity to the TraD mobilization proteins encoded by plasmids F and R100 from Escherichia coli. Nucleotide sequence analysis of one NTBF pattern III strain (strain I-1345) revealed that bfmB and bfmC are adjacent to each other and separated by a 16-bp GC-rich sequence. Comparison of this sequence with the appropriate sequence of ETBF strain 86-5443-2-2 showed that in this ETBF strain the 16-bp sequence is replaced by the BfPAI. This result defined the BfPAI as being 6,036 bp in length and its precise integration site as being between the bfmB and bfmC stop codons. The G+C content of the BfPAI (35%) and the flanking DNA (47 to 50%) differ greatly from that reported for the B. fragilis chromosome (42%), suggesting that the BfPAI and its flanking region are two distinct genetic elements originating from very different organisms. ETBF strains may have evolved by horizontal transfer of these two genetic elements into a pattern II NTBF strain.
Figures



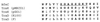

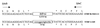
Similar articles
-
Characterization of the Bacteroides fragilis pathogenicity island in human blood culture isolates.Anaerobe. 2006 Feb;12(1):17-22. doi: 10.1016/j.anaerobe.2005.06.005. Epub 2005 Aug 15. Anaerobe. 2006. PMID: 16701608
-
Enterotoxigenic and non-enterotoxigenic Bacteroides fragilis from fecal microbiota of children.Braz J Microbiol. 2015 Oct-Dec;46(4):1141-5. doi: 10.1590/S1517-838246420140728. Epub 2015 Oct 27. Braz J Microbiol. 2015. PMID: 26691473 Free PMC article.
-
Modulation of bft expression by the Bacteroides fragilis pathogenicity island and its flanking region.Mol Microbiol. 2002 Aug;45(4):1067-77. doi: 10.1046/j.1365-2958.2002.03077.x. Mol Microbiol. 2002. PMID: 12180925
-
The toxins of Bacteroides fragilis.Toxicon. 2001 Nov;39(11):1737-46. doi: 10.1016/s0041-0101(01)00160-x. Toxicon. 2001. PMID: 11595636 Review.
-
Enterotoxigenic Bacteroides fragilis: a rogue among symbiotes.Clin Microbiol Rev. 2009 Apr;22(2):349-69, Table of Contents. doi: 10.1128/CMR.00053-08. Clin Microbiol Rev. 2009. PMID: 19366918 Free PMC article. Review.
Cited by
-
Enterotoxigenic Bacteroides fragilis.Curr Infect Dis Rep. 2008 May;10(2):99-104. doi: 10.1007/s11908-008-0018-7. Curr Infect Dis Rep. 2008. PMID: 18462582
-
Pathogenicity islands in bacterial pathogenesis.Clin Microbiol Rev. 2004 Jan;17(1):14-56. doi: 10.1128/CMR.17.1.14-56.2004. Clin Microbiol Rev. 2004. PMID: 14726454 Free PMC article. Review.
-
Structure, function and latency regulation of a bacterial enterotoxin potentially derived from a mammalian adamalysin/ADAM xenolog.Proc Natl Acad Sci U S A. 2011 Feb 1;108(5):1856-61. doi: 10.1073/pnas.1012173108. Epub 2011 Jan 13. Proc Natl Acad Sci U S A. 2011. PMID: 21233422 Free PMC article.
-
Structural and functional diversity of metalloproteinases encoded by the Bacteroides fragilis pathogenicity island.FEBS J. 2014 Jun;281(11):2487-502. doi: 10.1111/febs.12804. Epub 2014 Apr 22. FEBS J. 2014. PMID: 24698179 Free PMC article.
-
Bile acid fitness determinants of a Bacteroides fragilis isolate from a human pouchitis patient.mBio. 2024 Jan 16;15(1):e0283023. doi: 10.1128/mbio.02830-23. Epub 2023 Dec 8. mBio. 2024. PMID: 38063424 Free PMC article.
References
-
- Chung, G.-T., A. A. Franco, and C. L. Sears. Unpublished data.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

