Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments
- PMID: 10543824
- PMCID: PMC91682
- DOI: 10.1128/AEM.65.11.5066-5074.1999
Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments
Abstract
The 16S rRNA and pmoA genes from natural populations of methane-oxidizing bacteria (methanotrophs) were PCR amplified from total community DNA extracted from Lake Washington sediments obtained from the area where peak methane oxidation occurred. Clone libraries were constructed for each of the genes, and approximately 200 clones from each library were analyzed by using restriction fragment length polymorphism (RFLP) and the tetrameric restriction enzymes MspI, HaeIII, and HhaI. The PCR products were grouped based on their RFLP patterns, and representatives of each group were sequenced and analyzed. Studies of the 16S rRNA data obtained indicated that the existing primers did not reveal the total methanotrophic diversity present when these data were compared with pure-culture data obtained from the same environment. New primers specific for methanotrophs belonging to the genera Methylomonas, Methylosinus, and Methylocystis were developed and used to construct more complete clone libraries. Furthermore, a new primer was designed for one of the genes of the particulate methane monooxygenase in methanotrophs, pmoA. Phylogenetic analyses of both the 16S rRNA and pmoA gene sequences indicated that the new primers should detect these genes over the known diversity in methanotrophs. In addition to these findings, 16S rRNA data obtained in this study were combined with previously described phylogenetic data in order to identify operational taxonomic units that can be used to identify methanotrophs at the genus level.
Figures
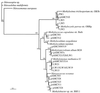
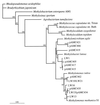
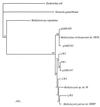
References
-
- Auman, A., and M. E. Lidstrom. Unpublished data.
-
- Auman, A., S. Stolyar, and M. E. Lidstrom. Unpublished data.
-
- Bender M, Conrad R. Methane oxidation activity in various soils and freshwater sediments: occurrence, characteristics, vertical profiles and distribution on grain size fractions. J Geophys Res. 1994;99:16531–16540.
-
- Bodrossy L, Holmes E M, Holmes A J, Kovacs K L, Murrell J C. Analysis of 16S rRNA and methane monooxygenase gene sequences reveals a novel group of thermotolerant and thermophilic methanotrophs, Methylocaldum gen. nov. Arch Microbiol. 1997;168:493–503. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

